Signals One™ - What's New
Signals One™ is a complete, integrated data capture, processing & analysis SaaS software for scientific research. Capture, manage, and share experiment data, notes and research findings in a secure and accessible environment built on the foundation of the Signals Platform.
Check out the latest improvements:
Releases By Year
Revvity Signals™ One, including Signals Notebook, Update
What’s New
We are delighted to release our new version of Signals, 26.4.
The following improvements are available for users, administrators and developers on the Signals platform. Certain features may only be available with appropriate licensing and/or with enablement by an administrator.
- Fundamentals
- Ontologies integration allows terms to be used across experiments, materials, tables, and more available to all systems
- Ontology integration now supported in Synergy, including Collaboration, Idea, Work Order, and Synergy Experiments Templates, as well as Designs and Reference Table fields available to all systems
- CSV import handling for ontology terms
- Ontology support for plate templates
- External Ontology Sources now support SciBite CENtree
- Audit trail displays highlighted granular changes for text elements (beta)
- AI
- Prompt-to-filter (beta)*
- Data Workflows & Analytics*
- Data Assembly continuous viewer
- Median-based robust Normalizations
- Additional Quality Control metrics
- SAR & Decisioning*
- Ontology support on Inventa Analysis templates
- New ChemCharts Biopolymer data functions
- ChemCharts Table usability improvements
- Synergy†
- Collaboration setting allows Sponsor admin to choose whether to use Material IDs or aliased ExIDs with CROs
- Search for Synergy Experiments based on Work Order
- System Snippets are available for CRO users
- Work Tasks in Work Orders (beta)
- Chemistry
- Multi-strand and branched complex biopolymers now supported
- Upload of rdf file types to External Enumerator
- Tables
- Definition and display of Scatter Plots on Admin Defined Tables available to all systems*
- End user adjustment of Scatter Plots available to all systems*
- Export of image from Scatter Plots available to all systems*
- Highlight selected rows or points in the Scatter Plot and Table available to all systems*
- Definition and display of Bar Charts on Admin Defined Tables*
- Define and display Size By and Transparency on Scatter Plot to enable Bubble Charts on Admin Defined Tables*
- Inventory
- Enable container ID and barcode selection directly within Reagent tables
- Support Inventory Decrement at Sub‑Experiment Level in Parallel Experiments
- Display container details in Samples table when Material ID is present
- Multi Select property display in the Container search table
- InventoryGo‡
- Flexible reconciliation workflows introduced: Batch vs. Individual scanning options
- Administration
- Migration for legacy Data Factory projects
- Spotfire for Signals Online Access privilege available
- Admin Defined Dashboards
- Audit trail displays highlighted granular changes for admin-defined tables (beta)
- Integrations & Interfaces
- Ontology API Support
- Salt Stripping Data for Chemical Drawings fetch API
- Entity properties on Bulk Export API
- Update Hierarchical Tables Headers API
- External Enumerator via API
- Bulk create Samples via Summary Table via API
* Requires Signals One license
† Requires Signals Synergy license
‡ Requires InventoryGo license
We also fixed several small bugs in this release. Details of the enhancements are described below.
Administrators are recommended to subscribe to the channels within our support news site found at https://support.revvitysignals.com/hc/en-us/categories/360004446171-Support-News which contains more information about releases and other pertinent product information.
Data Factory administrators should be aware that the Bulk Delete option from the Inventa Dashboard is now available regardless of the “Enable system-generated Unique Result IDs in Spotfire workflows” setting. However, Using Bulk Delete still requires explicit mapping of a Unique Result ID.
Data Factory Administrators should note that legacy Data Factory projects will be fully deprecated in a future release. A straightforward migration path is now available to support this transition. A specific deadline has not yet been scheduled.
Data Factory administrators should also note that the Advanced Security policies in Inventa apply an “AND” logic across all user groups, restricting access if any groups lack permissions. This behavior differs from the standard Signals security model and will be fixed to align with the standard in a future release.
This content is anticipated for release on our Production E3 environments, and for Private Cloud customers on our deferred release schedule, in November 2026.
Further Details
The following improvements are available for users, administrators and developers on the Signals platform. Certain features may only be available with appropriate licensing and/or with enablement by an administrator.
Fundamentals
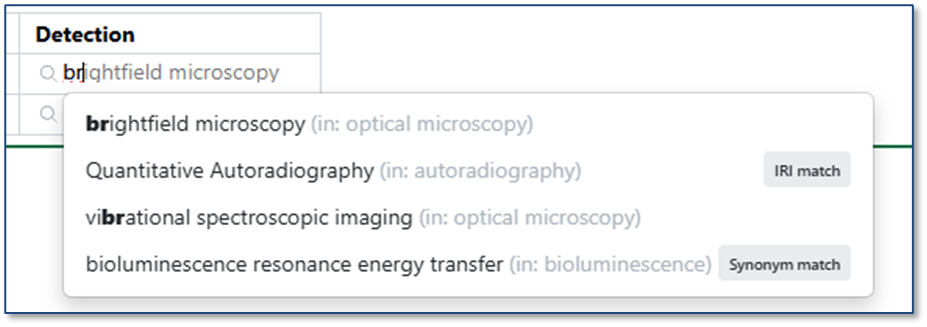
Ontology support is now available for all systems. Administrators may connect to a ontology provider (BioPortal, OLS, or SciBite CENtree) as an External Ontology. Once a specific ontology from that source is connected, Administrators can create a new type of Attribute, Ontology Lists, which contain a subset of terms from the ontology. This Ontology List can be used in templates and properties across signals, including Experiments, Materials, Tables, Samples, Admin Defined Objects, Plates, Inventa Analyses (for Signals One), Synergy Experiments and Design/Reference Tables, and more.
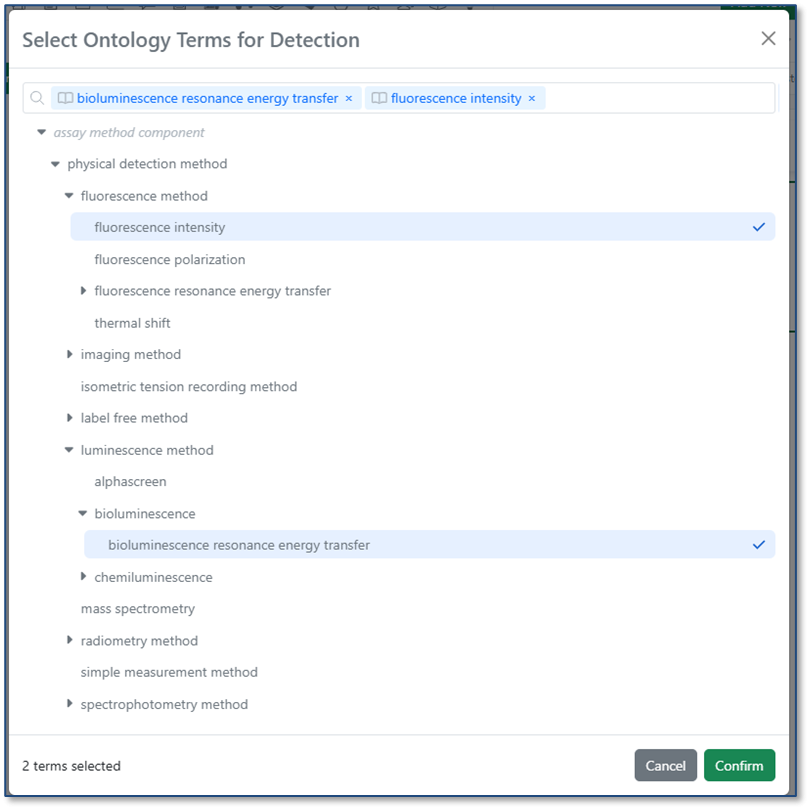
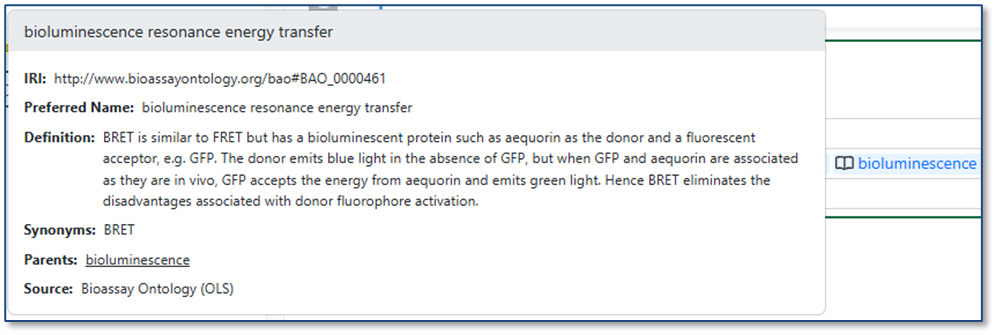
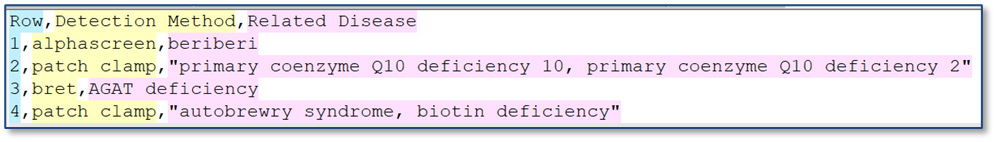
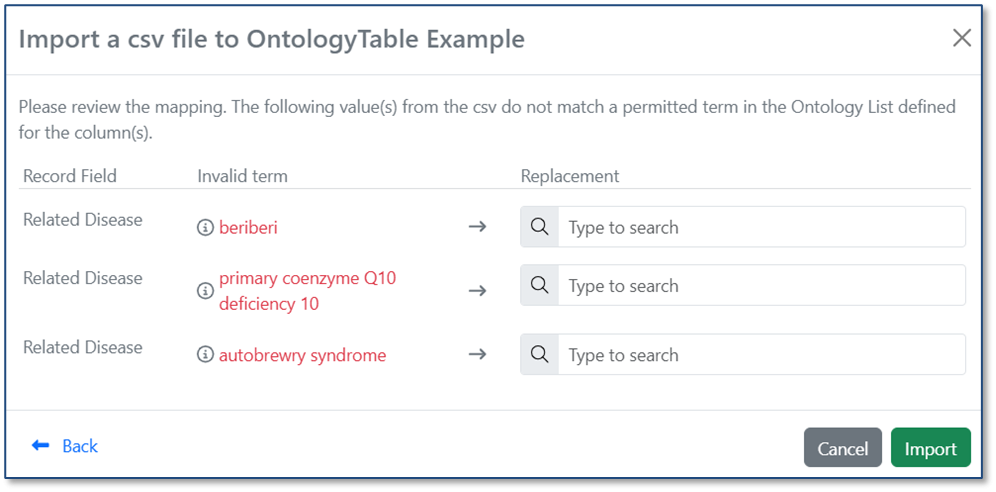
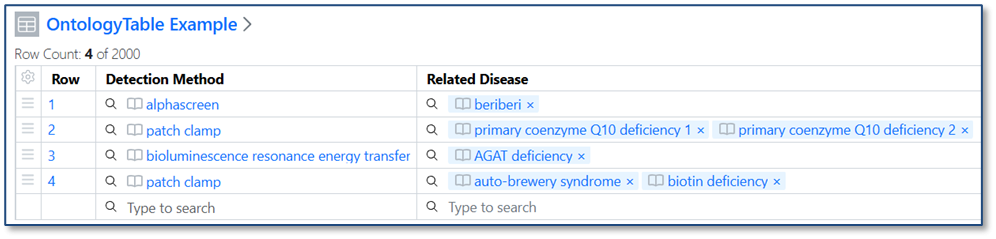
When you import a CSV into a table, ontology values are now validated against the Ontology List for that field. If a value in the CSV matches a term’s preferred name or any synonym, it is automatically mapped to the correct ontology term. If no matching term is found, users can resolve the value during import by selecting the appropriate term from the Ontology List.
Data Workflows & Analytics
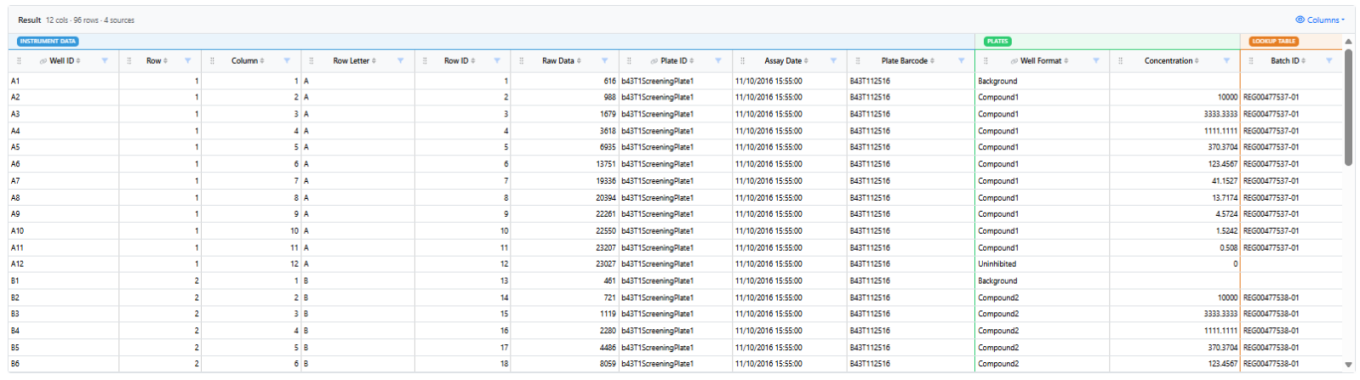
In Signals One, all In Vitro Analysis users will discover that in the Data Assembly step, the 5-rows static preview has been replaced by an interactive viewer with continuous scroll. Numerical and categorical columns can now be filtered, sorted, reordered and hidden to flexibly slice and dice the data that will be used in the Analytics steps. Visual cues indicate the source data of the columns, and which columns were used during the matching process.
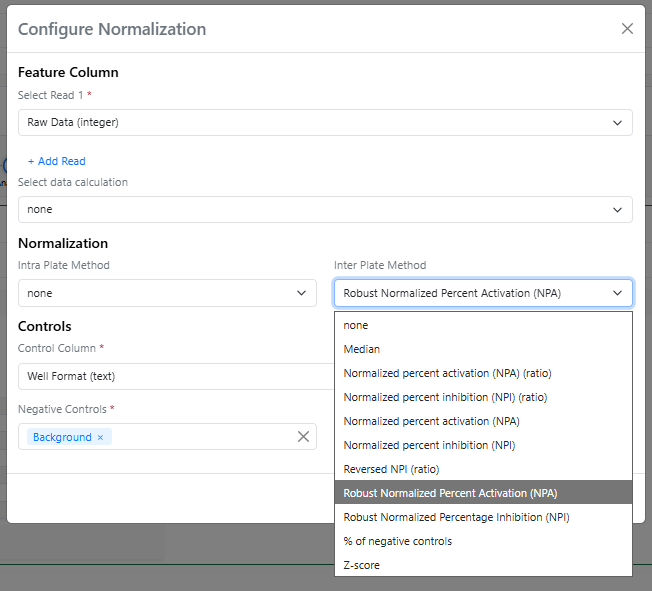
In the Analytics steps of the In Vitro Analysis, a couple of new normalization methods, using the median instead of the average, can now be found in the Inter Plate Method menu, both with the “Robust” prefix.
The Robust Normalized Percent Activation is calculated using the expression: “(value - mean(n)) / (mean(p) - mean(n)) * 100” while the Robust Normalized Percent Inhibition is calculated using the expression: “(mean(p) – value) / (mean(p) - mean(n)) * 100” (where p and n are the values for positive and negative controls). Median-based normalizations are called "Robust" because they are not heavily influenced by extreme outliers.
Finally, the mean and standard deviation of both positive and negative controls are calculated and added to the Normalization results.
SAR & Decisioning
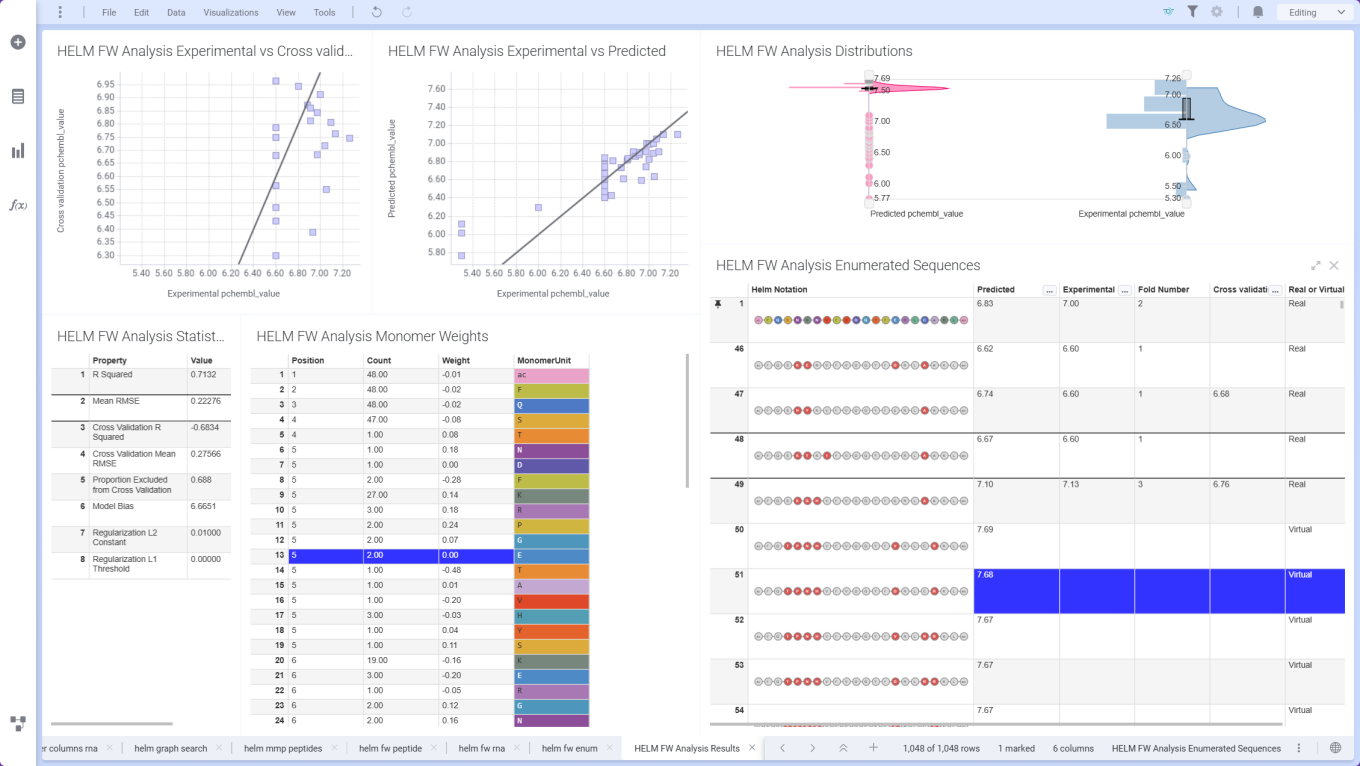
ChemCharts Data Functions introduce Free‑Wilson analysis for HELM sequences, using aligned sequence and property data to quantify position‑specific monomer contributions. Combined with the Free‑Wilson Enumerator, users can design and prioritize novel HELM sequences based on predicted performance.
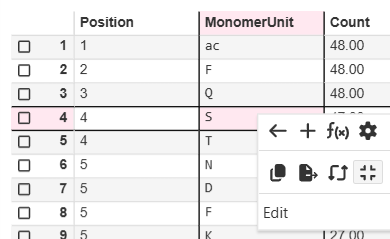
ChemCharts Table Visualizations now support a crosshair, which once activated highlights the cell, row number, and column header when moving the mouse.
Synergy
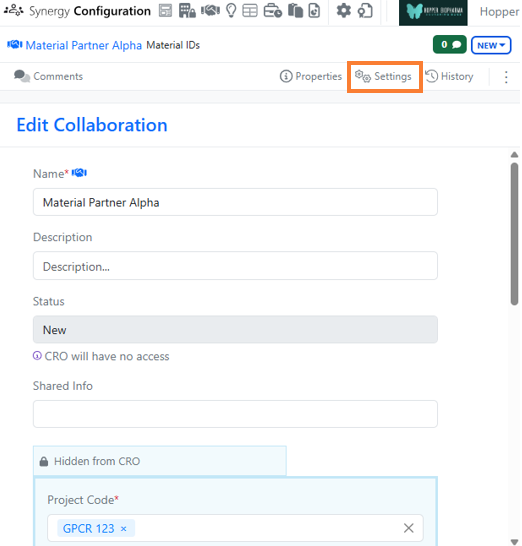
Synergy now allows the administrator to choose whether to use aliased ExIDs or Material IDs with a CRO on a Collaboration-by-Collaboration basis. An administrator with the privilege Manage Collaborations has access to a new Collaborations Settings menu on a Collaboration where the administrator can choose between aliased ExIDs or Material IDs. The default is ExIDs to support the current behavior. This setting can only be changed while the Collaboration is New (before being sent to the CRO) and no child Work Orders are yet created.
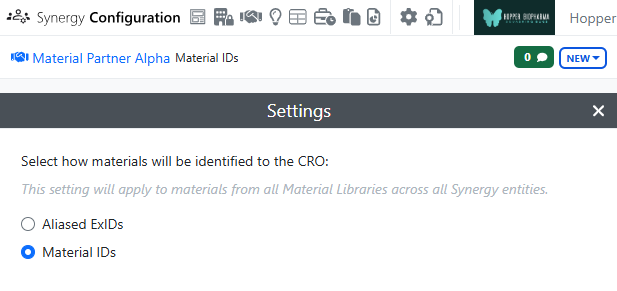
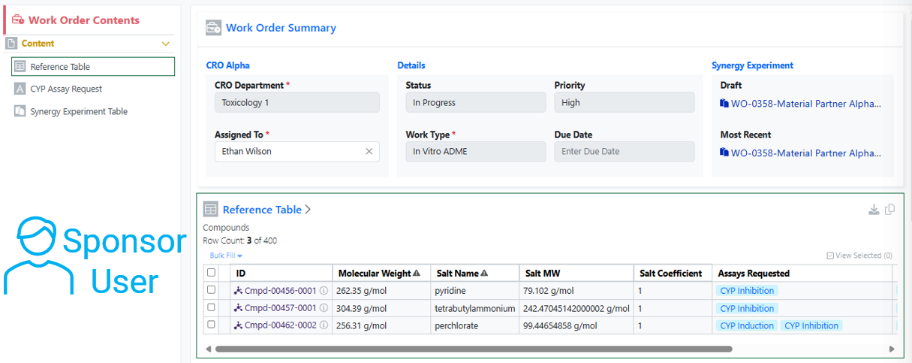
If the Collaboration is set to Materials IDs, materials shared via the Reference Table will display with their Material IDs to both the CRO and Sponsor. The Sponsor will not have a toggle to switch between the IDs, indicating that the Material ID will be shared with the CRO.
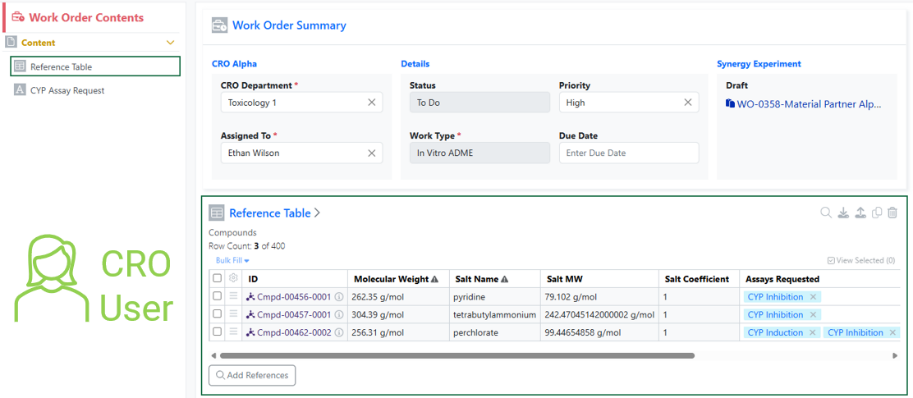
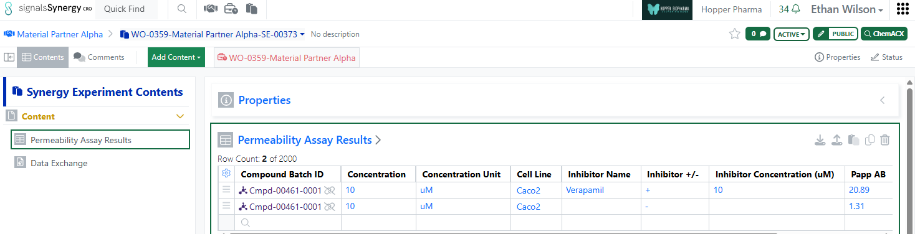
The use of Material IDs persists throughout the Collaboration in all child Work Orders and Synergy Experiments. For example, when a CRO user populates an internal reference column in an admin-defined table or when the Sponsor registers a Sample in a Samples table, the column will display the Material ID for both the Sponsor and CRO.
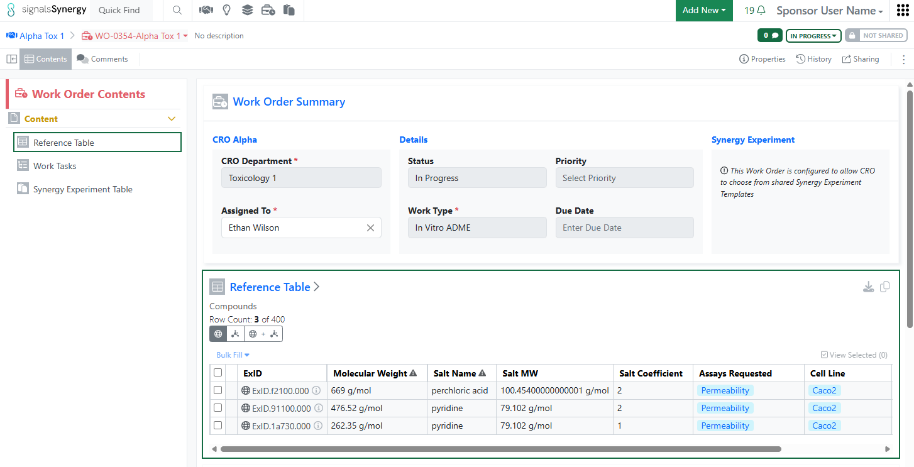
If the Collaboration is set to ExIDs (default), materials shared via the Reference Table will display with their ExIDs to the CRO, and the Sponsor will have a toggle to switch between the ExIDs, Material IDs, or both (existing behavior).
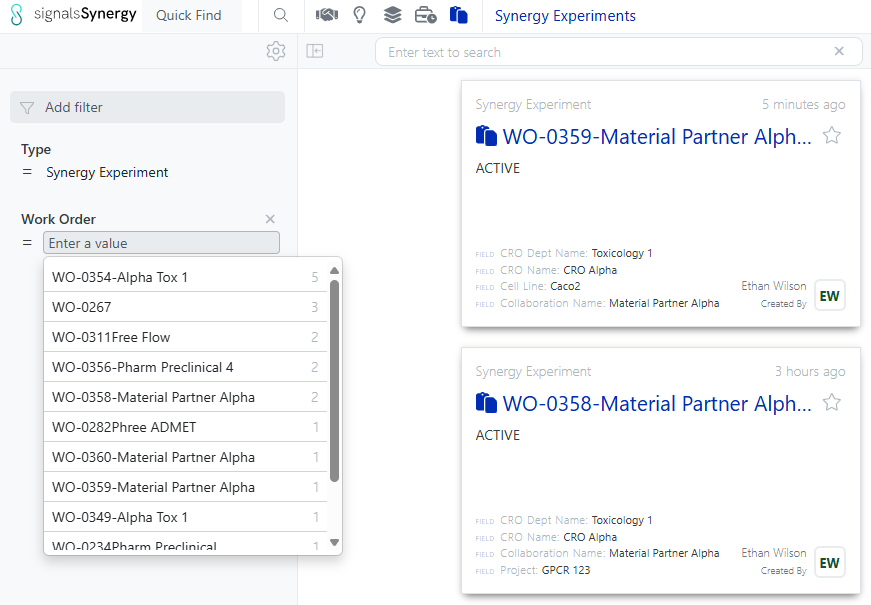
Both Sponsor and CRO users can now search for Synergy Experiments by the associated Work Order ID. The Work Order filter can be added in Advanced Search, typically alongside Type = Synergy Experiment, or to the Synergy Experiments Smart Folder.
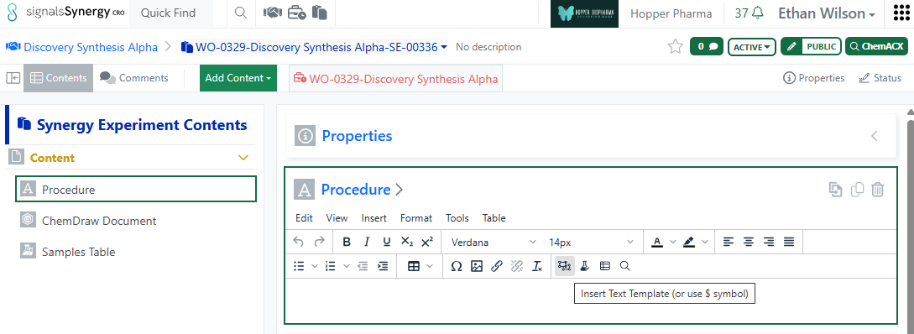
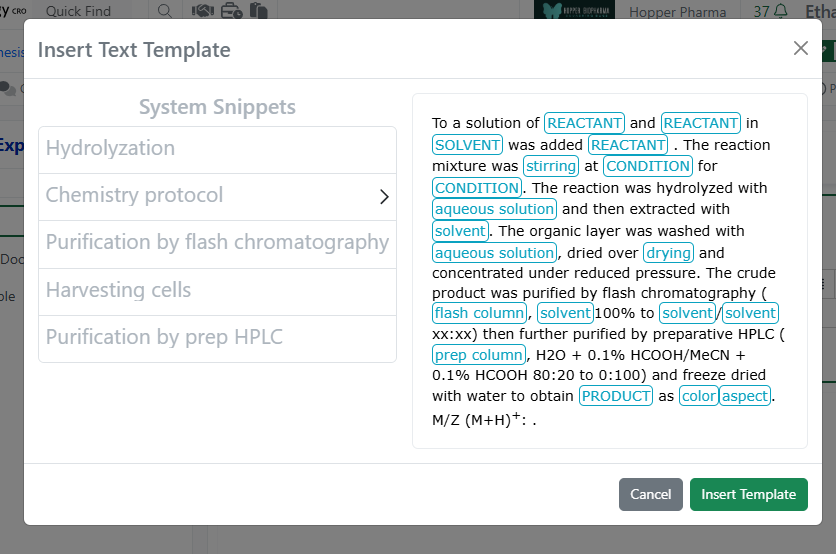
Finally, CRO users can now access and insert System Snippets using the Insert Text Template on any Text element. Note, CRO users cannot save text templates to avoid cluttering the template options for all users.
Chemistry
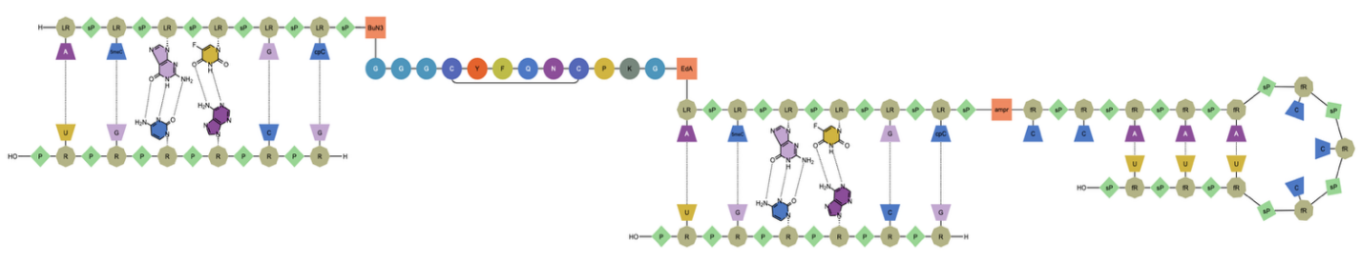
The ChemDraw editor now supports rendering and cleanup of multi-strand and branched complex biopolymers, making these structures easier to display in a clear, aligned, and readable layout.
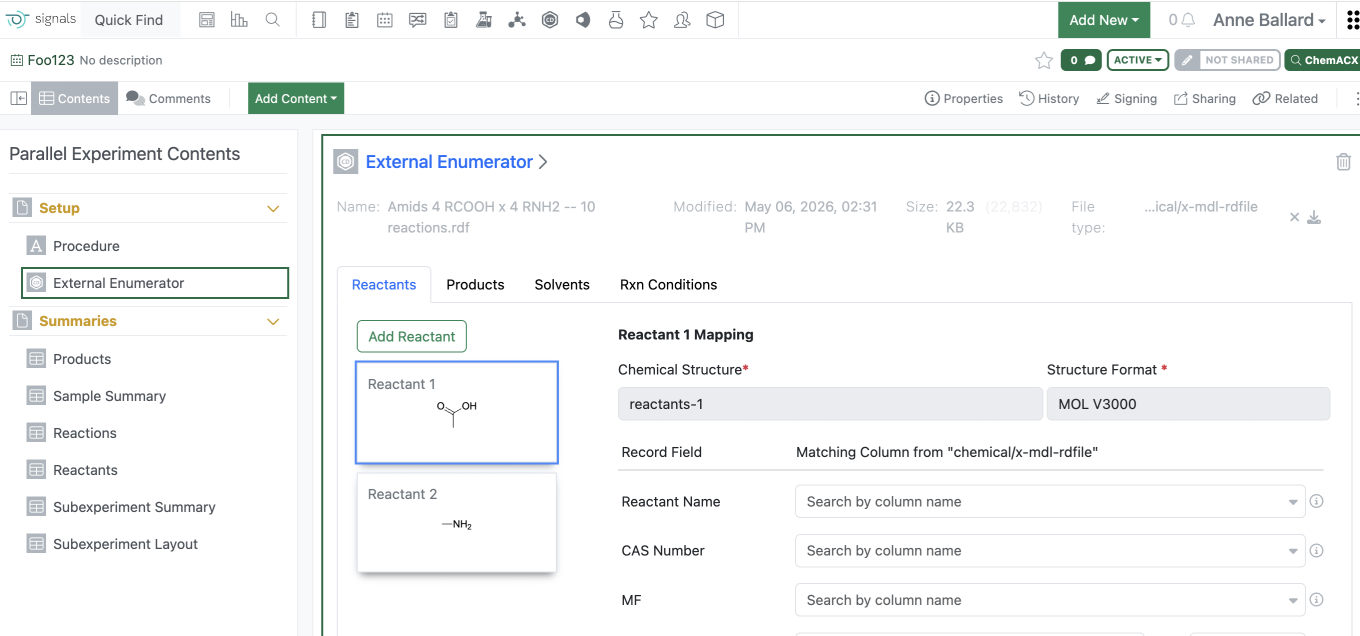
Users are now able to load an RDF type file into the External Enumerator for Parallel Experiment enumeration. Fields within the rdf can be mapped into the external enumerator. Reactant order is defined by the rdf file. Solvent and Rxn Conditions are not commonly included in the rdf file and therefore will need to be added manually.
Additional reactants or products can be added manually similarly to what can currently be managed.
Tables
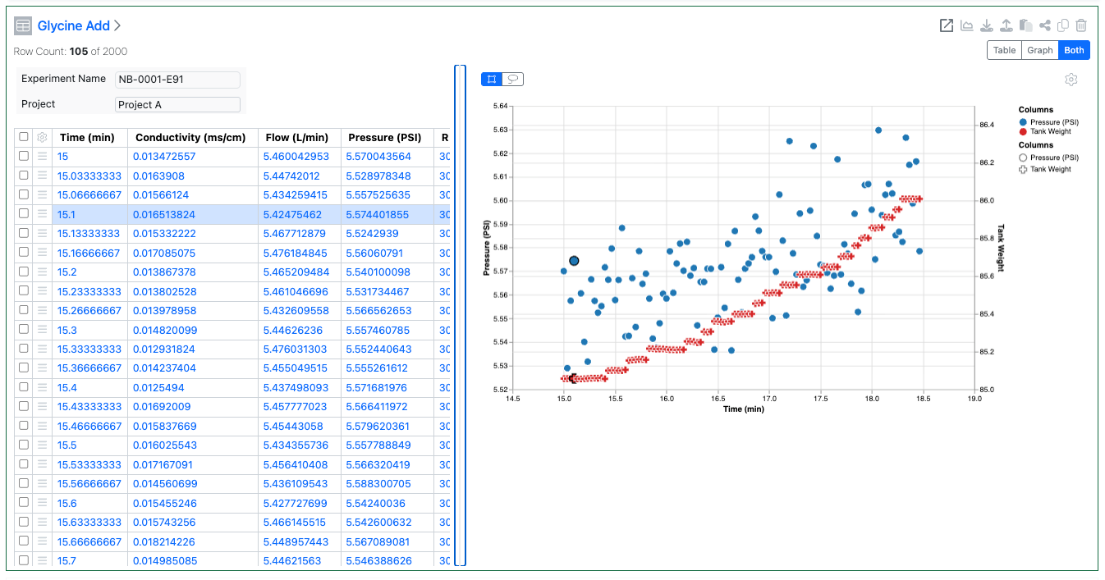
Scatter Plots are now available to all tenants. End users require a Signals One license to view and edit charts in ADTs.
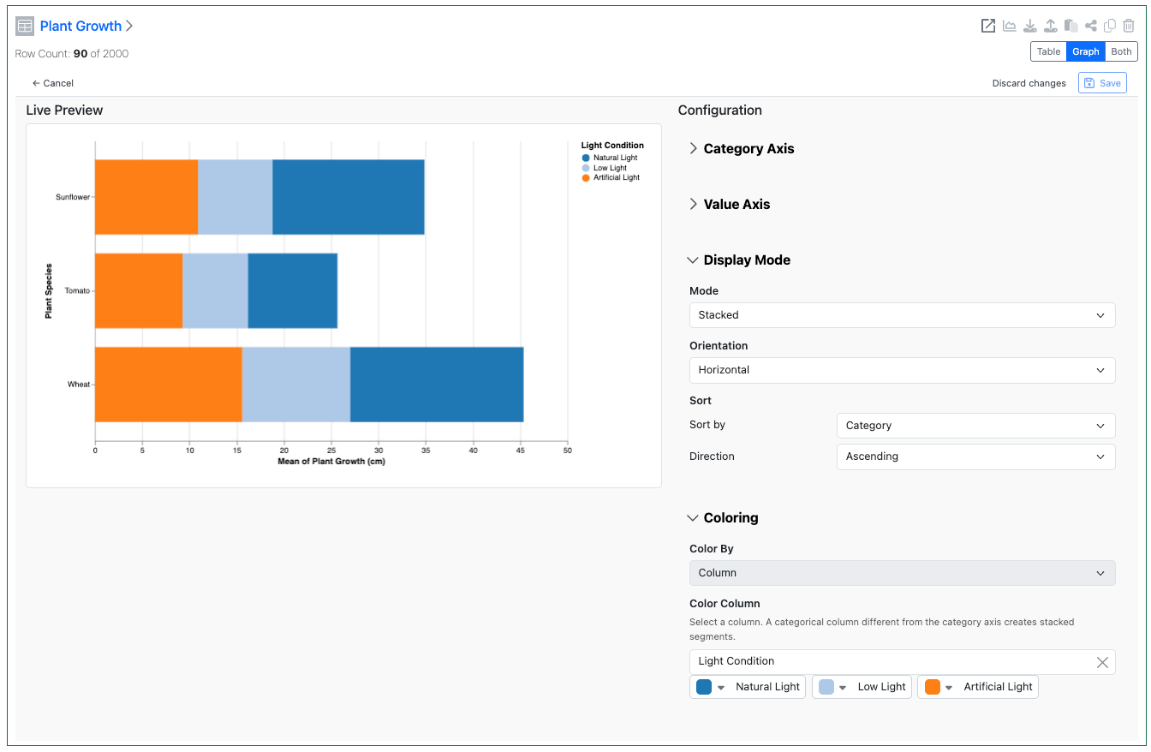
Configuration Administrators can now choose to define a Bar Chart on Admin Defined Tables. Similar to the definition of Scatter Plots, administrators can define the Bar Chart on the Table template including setting properties for the value and categorical axis. The chart can be a simple, stacked or 100% stacked bar chart and can be displayed vertically or horizontally. The admin can also choose if the end user is able to adjust the settings for the chart.
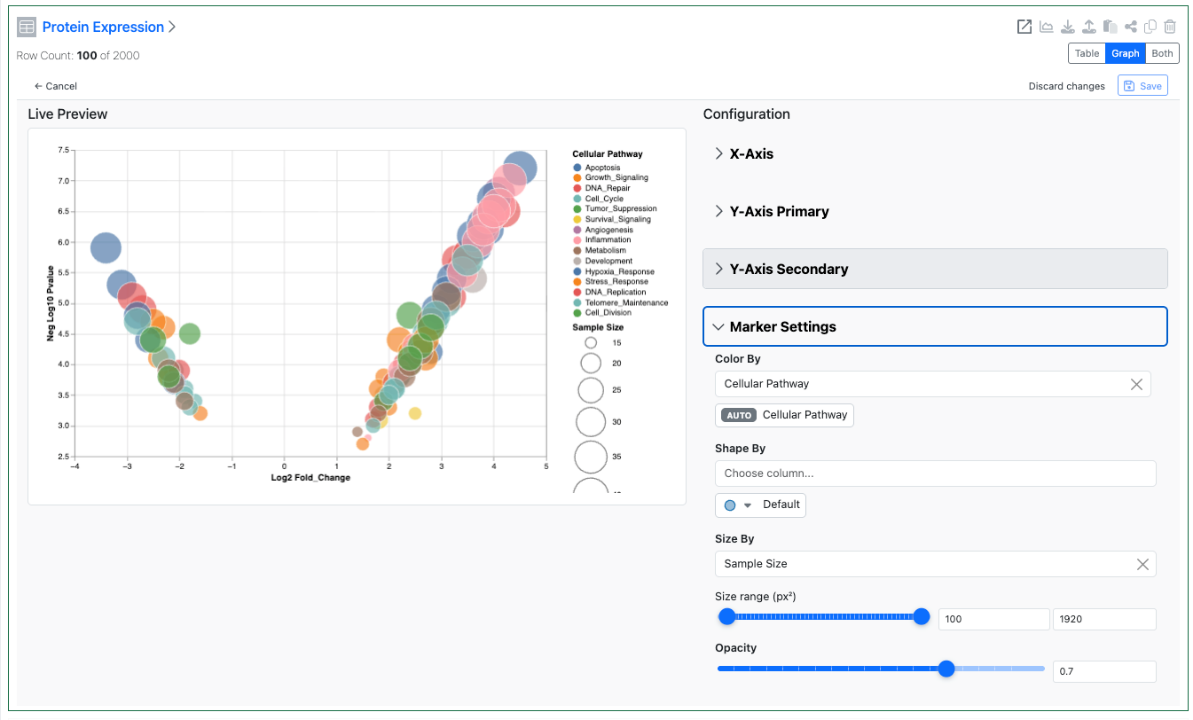
Configuration administrators can also now define a Size By and an Opacity setting for Scatter Charts. These two settings allow for the creation of Bubble Charts using the Scatter Chart type. If the end user is allowed to change chart settings these options are also available directly within the Admin Defined Table.
Inventory
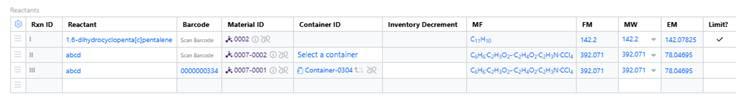
Users with the right permissions can now assign or reassign containers in cloned or pre-defined experiments, even after container details are cleared, enabling more flexible and accurate inventory workflows in Reagent table.
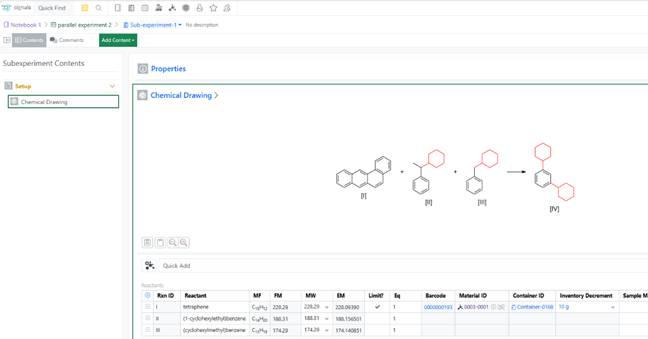
Inventory Decrement can now be entered at the sub‑experiment level, allowing users to accurately update container usage when working with parallel, cloned, or reagent‑specific sub‑experiments.
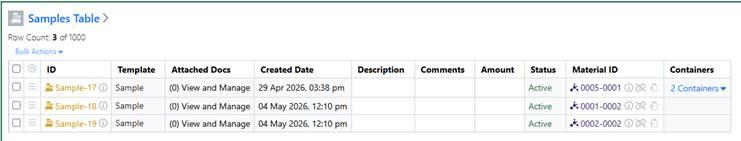
Container details are now accessible from the Samples table even when a Material ID is provided, ensuring consistent visibility across sample workflows.
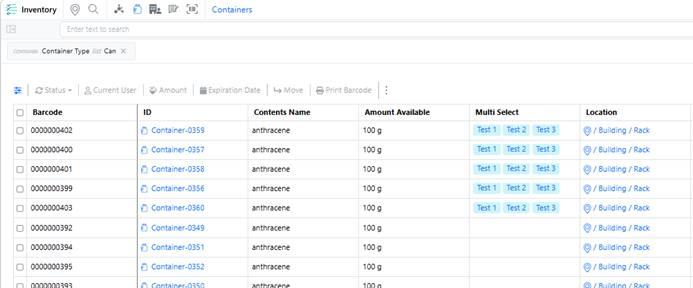
Container multi‑select properties are now available in the Container table, allowing users to view multi‑select container properties directly from the inventory view.
InventoryGo
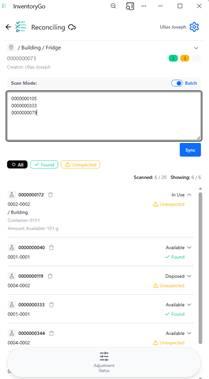
A new Scan List mode has been added to reconciliation, allowing users to scan containers in bulk and fetch results on demand, with a toggle to switch between batch and individual scanning for more reliable and faster workflows.
Administration
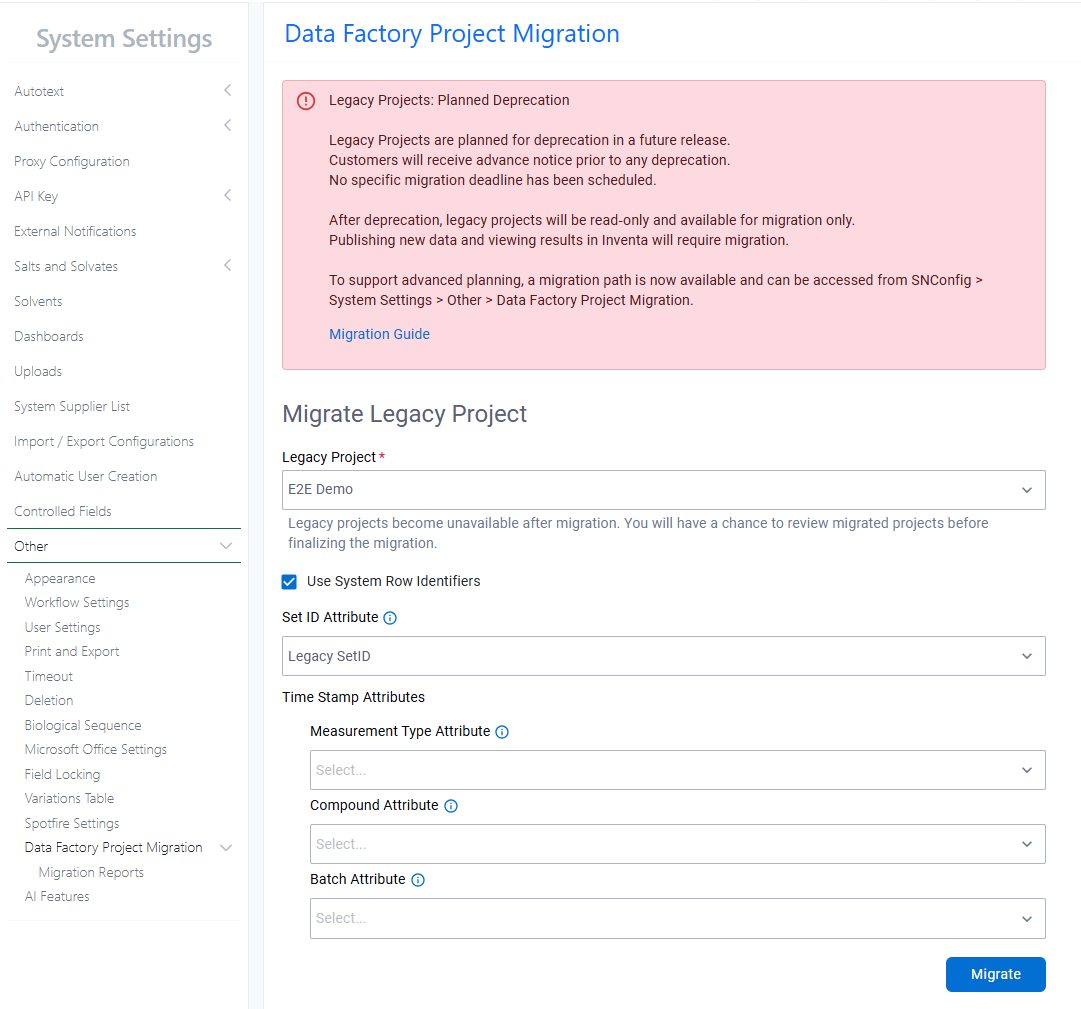
A migration path for legacy Data Factory projects is now available. Legacy Data Factory will be deprecated in a future release; administrators are encouraged to migrate to the latest Data Factory as soon as possible. The Legacy Project Migration tool is available from System Settings > Other. Before migrating, review the Migration Guide within the Signals Configuration Guide.
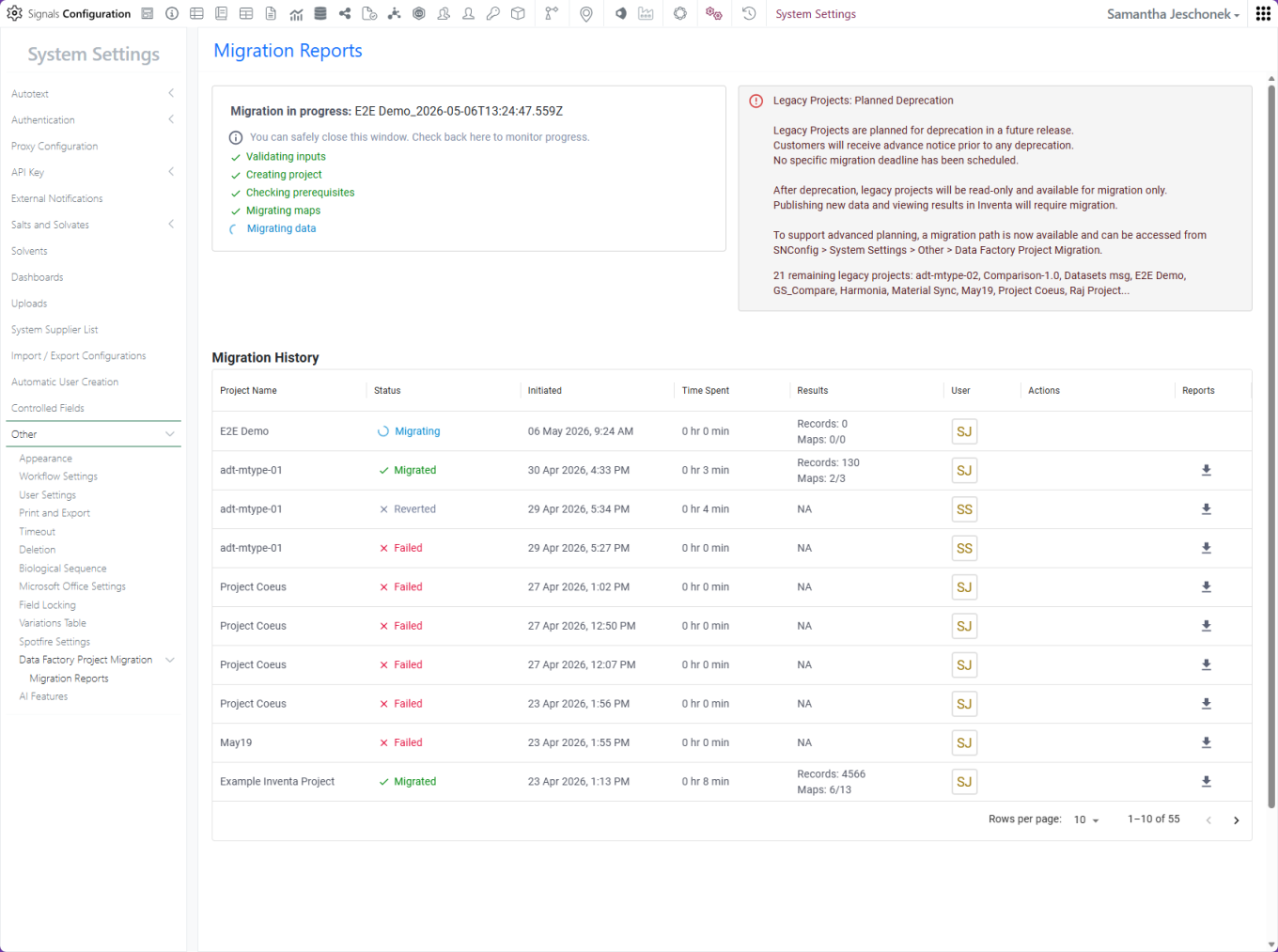
Migration status and history are captured in the Migration Report.
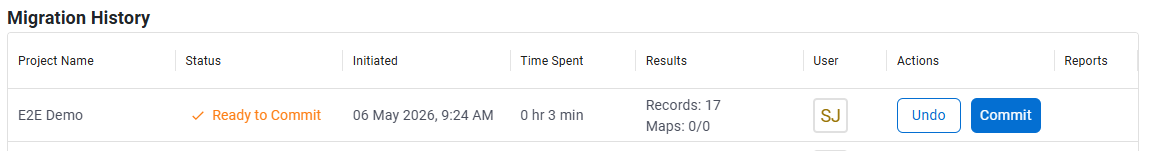
After a successful migration, administrators can review the migrated project before committing the changes.
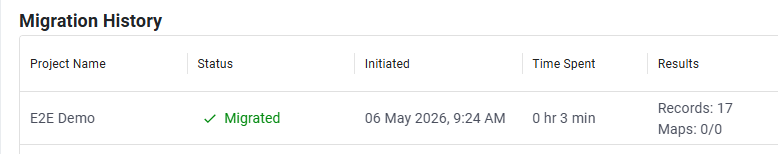
After you commit, the migrated project is ready to use and can be accessed from the Inventa Dashboard. The legacy project is no longer available for publication or data retrieval in Inventa.
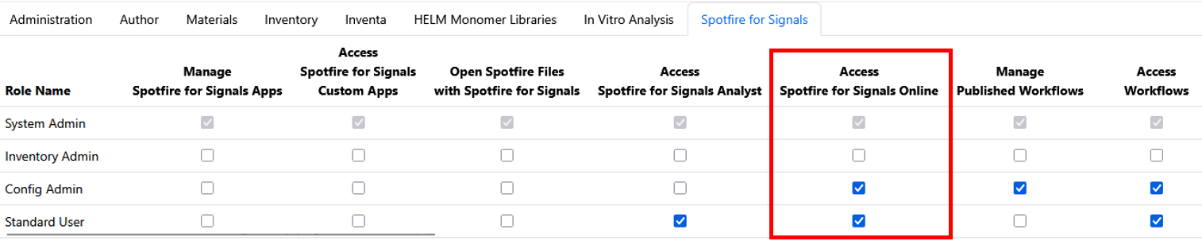
Spotfire for Signals Online Access privilege available to control which users can access and substantiate Spotfire for Signals Online. Users without this privilege will be limited to Spotfire for Signals Analyst use.
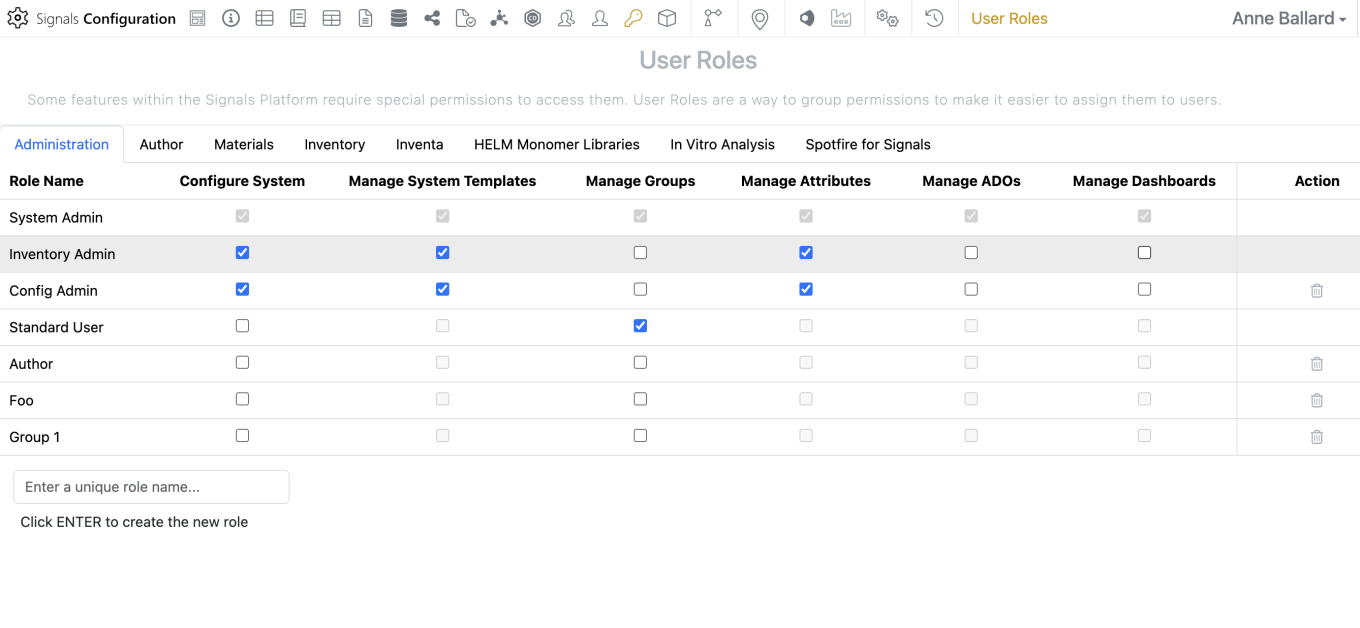
Administrators should now be able to create and share dashboards from the SN Config. To edit, share or create dashboards, a user must have the “Manage Dashboards” privilege and access to the SN Config as Admin Defined Dashboards can only be edited or created within the SN Config.
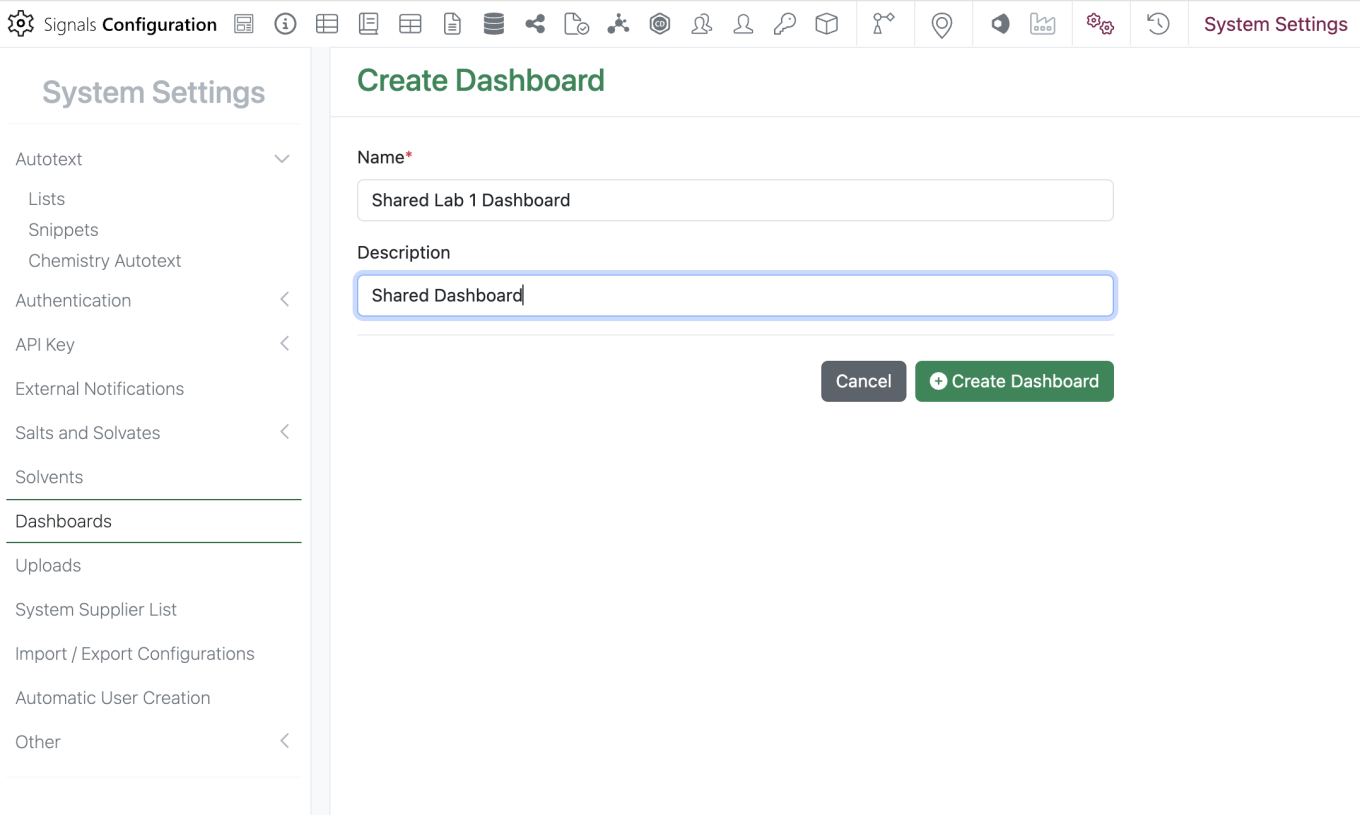
The new “Dashboards” section can be found in the System Settings. To create a dashboard, the admin should select the green “Create Dashboard” button then they are prompted to enter a dashboard name and description.
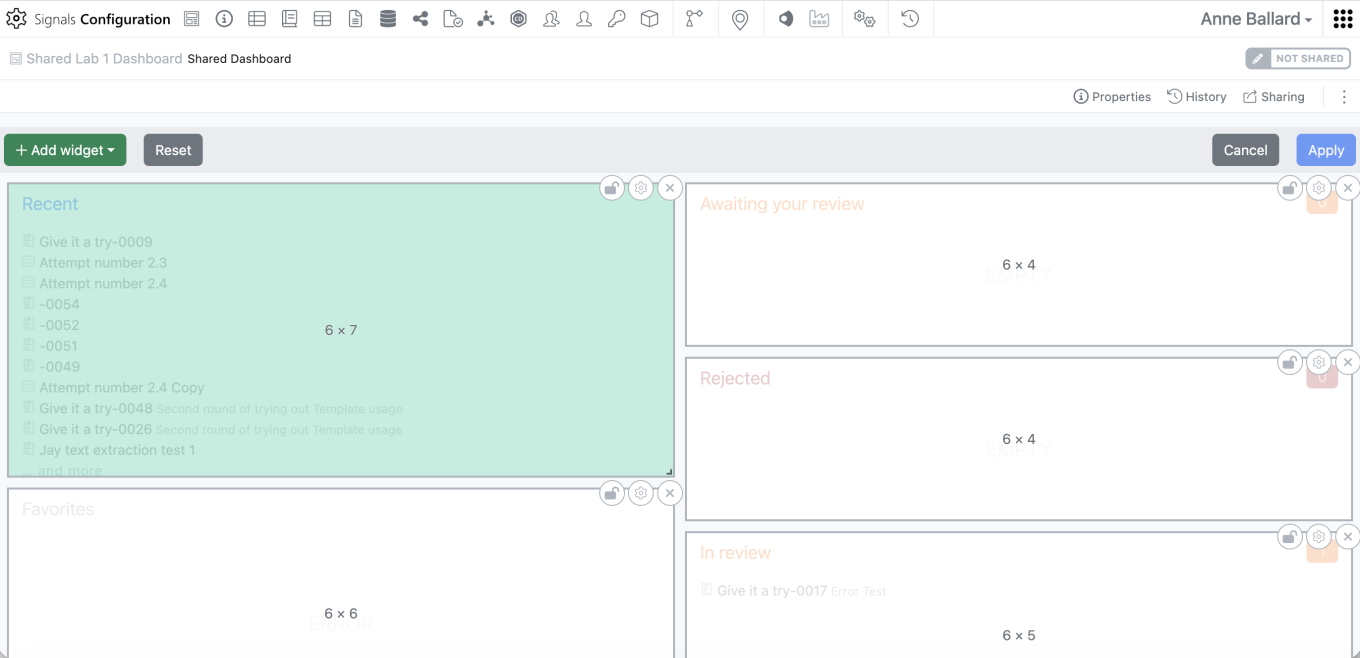
When the admin clicks on “Create Dashboard” they will be brought to the Dashboard edit screen. Here they can add, remove, edit, or resize widgets similarly to what can be done with a personal Dashboard.
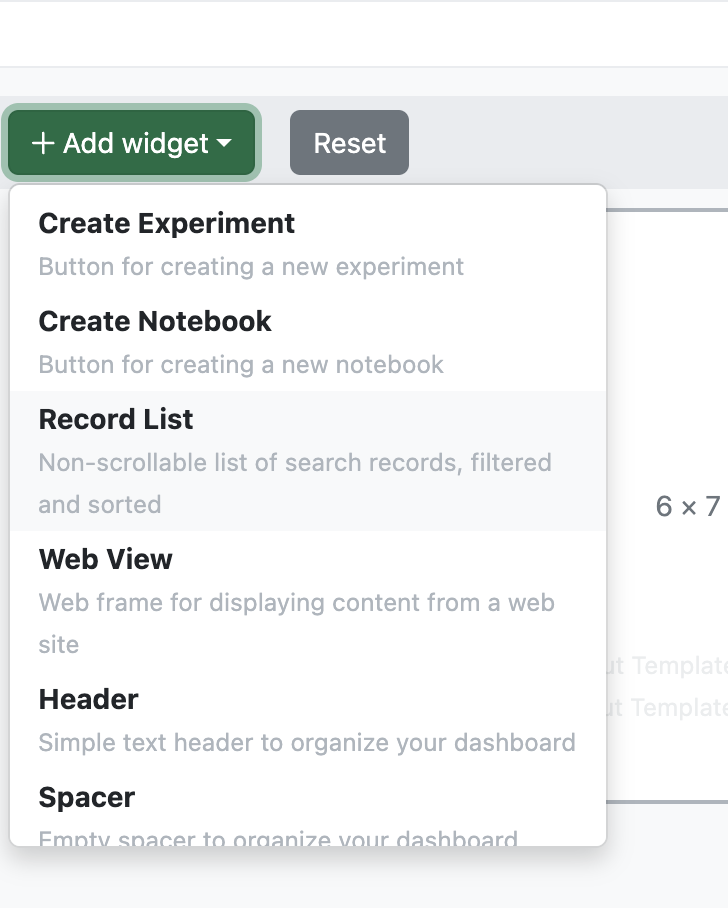
To add a new “Record List”, the most commonly utilized widget, the admin would select “Add Widget” > “Record List”. Under “List Source” the admin can select from a standard set of predefined lists or select “Define List” to create a new search list.
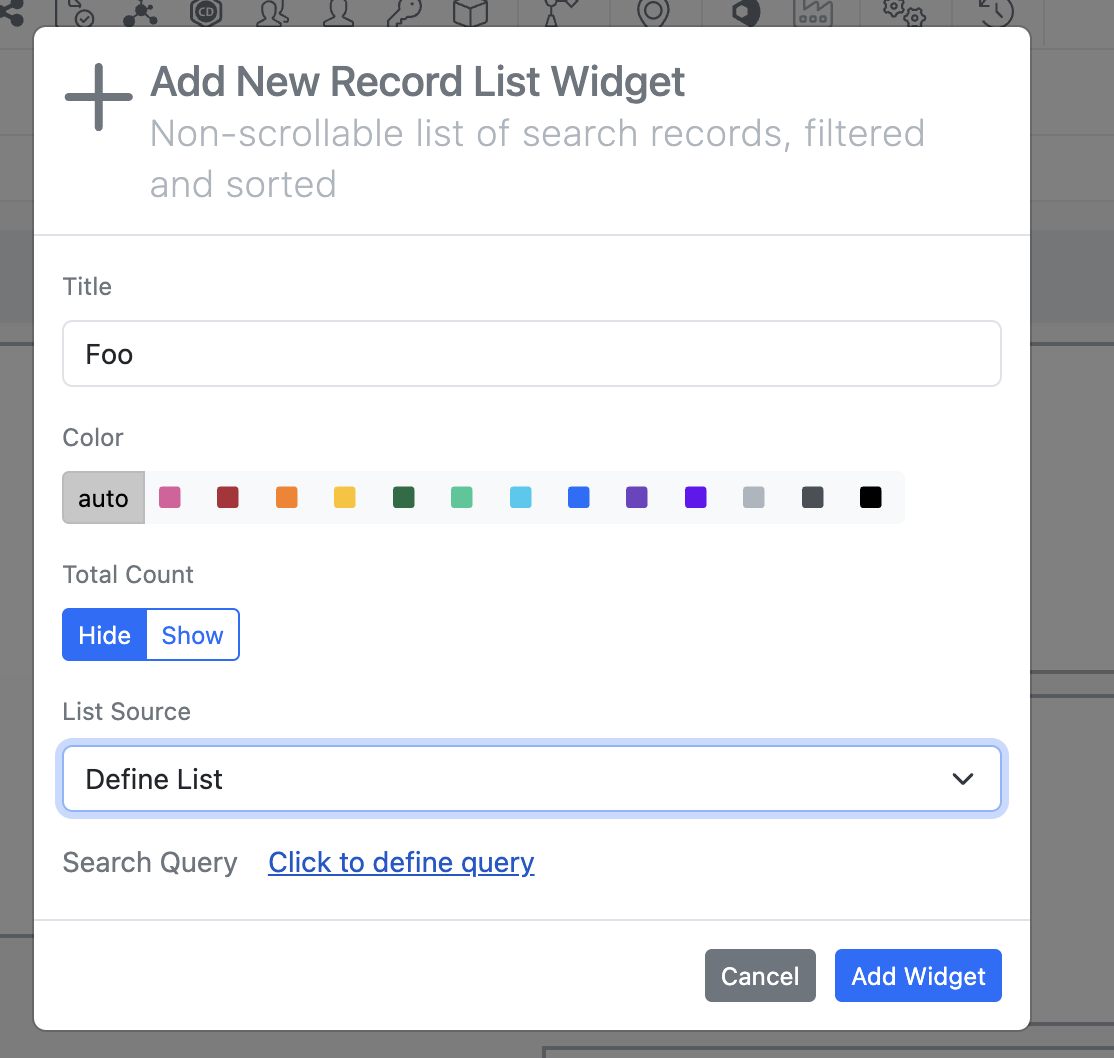
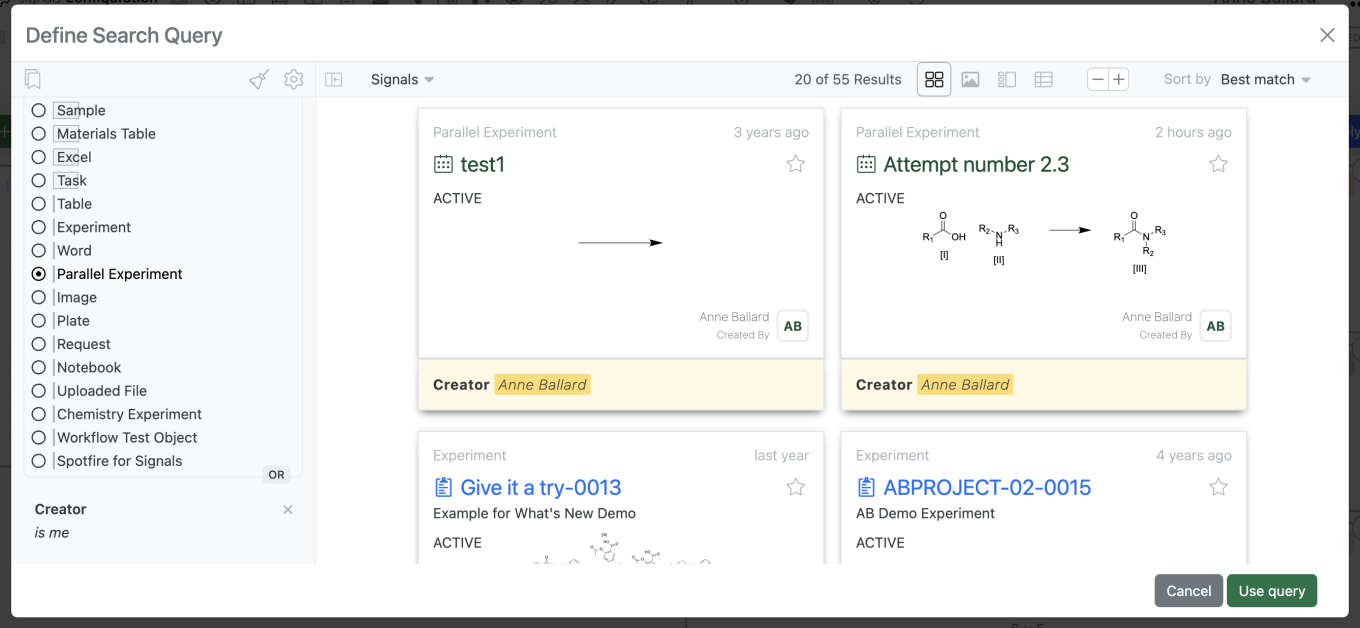
Selecting “Define List” > “Click to define query” brings up a search query builder the admin can utilize to define their search list.
Changes to the selected dashboard are saved by selecting the blue “Apply” in the top right corner.
The admin can then share the dashboard with other users or groups similarly to how sharing is managed in other elements.
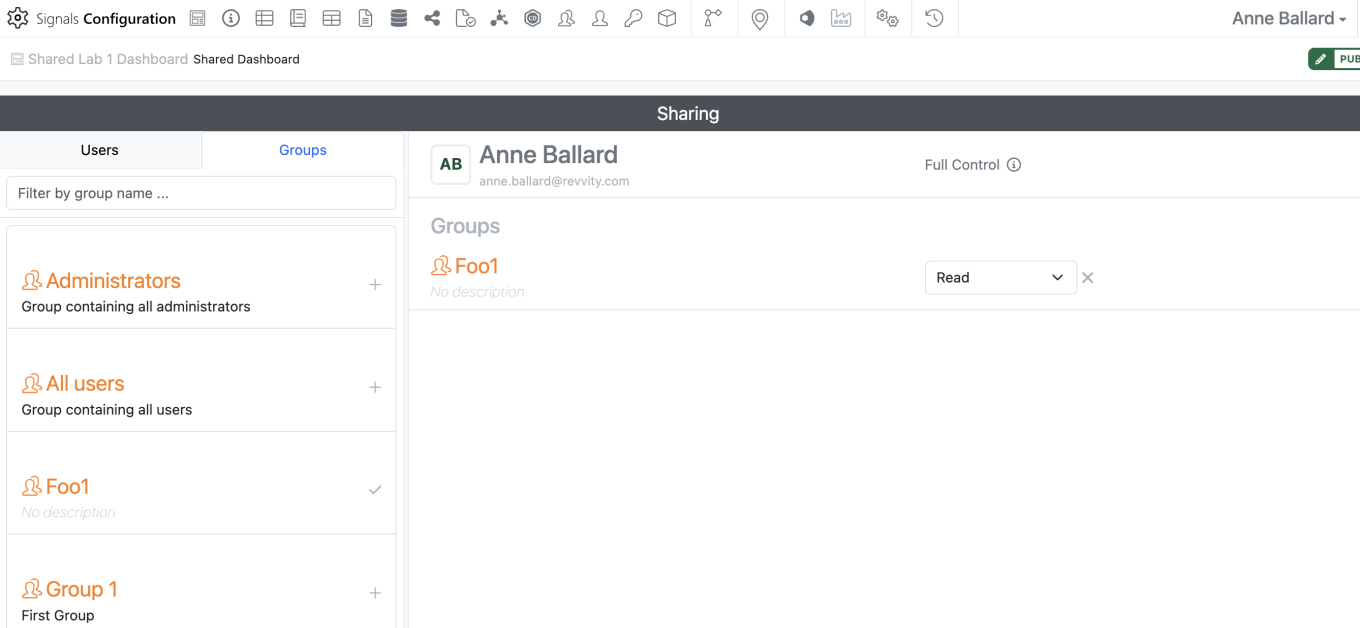
Dashboards are automatically shared with Read access. If a dashboard is shared with write access or full control to a user or group that does not have access to the SN Config, they will be unable to edit or share the dashboard as shared dashboards are Read Only outside of the SN Config.
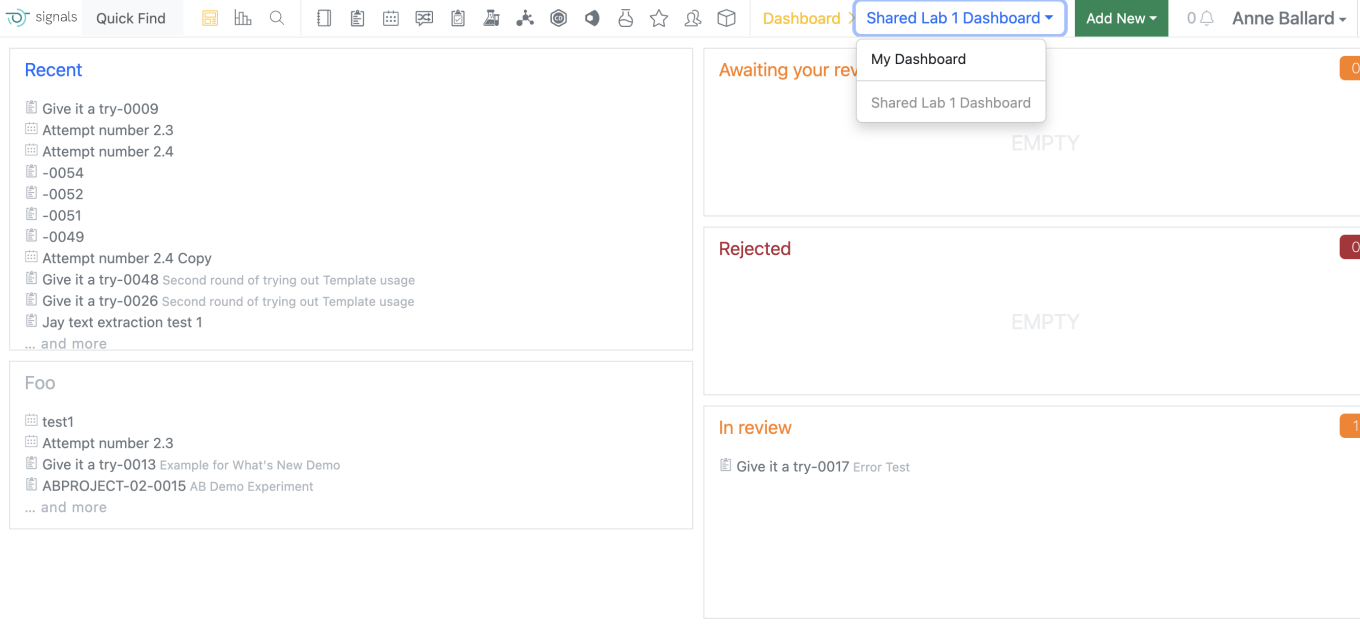
For end users, their current personal dashboard has now been named “My Dashboard” on the dashboard menu. Users are still able to edit their own “My Dashboard” as they were previously able to do. All shared dashboards appear below “My Dashboard”. If the user selects one of their shared dashboards this selection is sticky.
Integrations & APIs
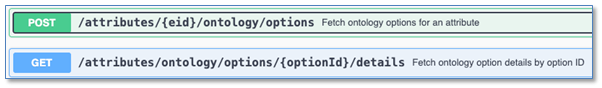
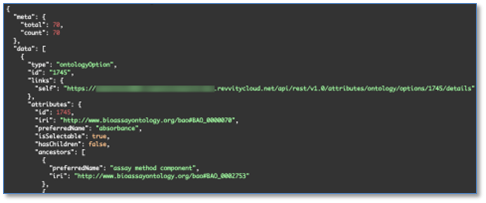
We have added API support for writing to Ontology List fields. These are enhancements to all APIs which write to entities that support Ontology List fields and properties. There are new APIs to fetch and explore Ontology Lists. The “id” values found here are used to set values on Ontology List fields and properties.
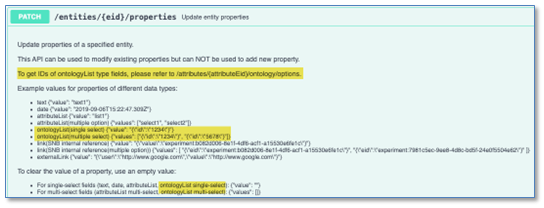

We have enhanced the stoichiometry fetch API to include salt stripping data for a Chemical Drawing. This is a new URL parameter (includeStrippedData) which when true will include the stripped structure and the salts that were stripped from the Chemical Drawing as data in the response.
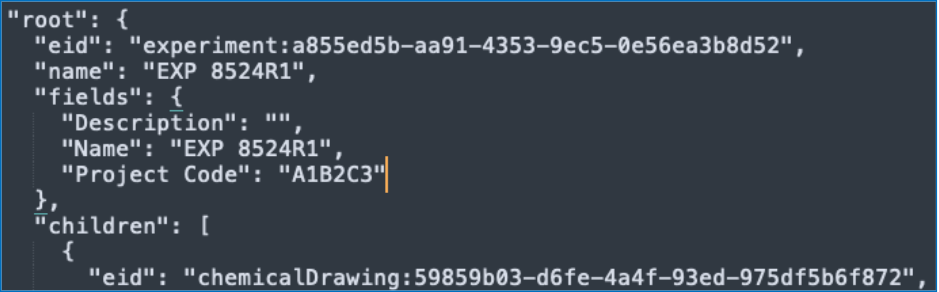
We have enhanced the API for bulk export of an entity to include the entities properties as part of the table of contents.

We have added a new API to allow patching of Hierarchical Table headers.
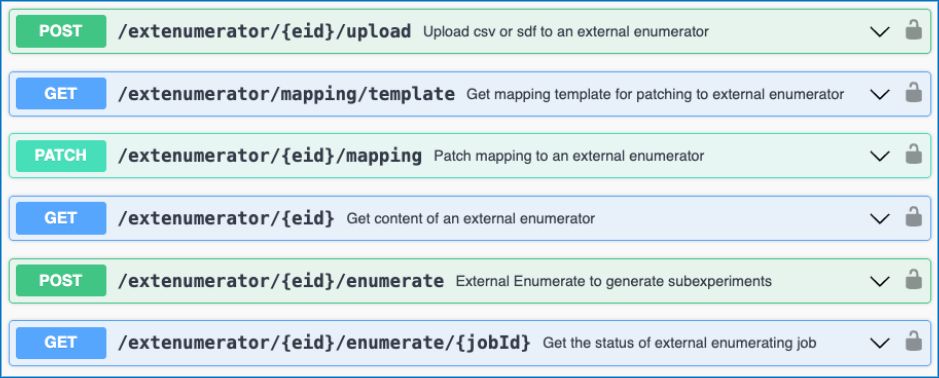
We have added new APIs to support the External Enumerator for Parallel Experiments. These APIs allow for usage of a csv or sdf file upload for use with the External Enumerator. There are APIs to fetch and update the mapping for the external enumerator. Lastly, there are APIs to kick off the enumeration and check its status.

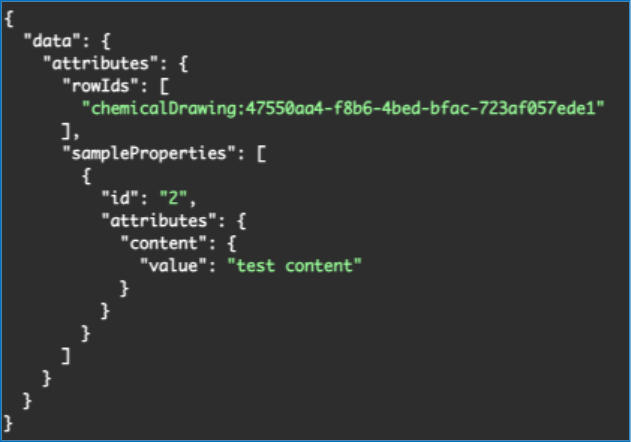
We have added new APIs to support the bulk creation of Samples in a Parallel Experiment Summary table. This is an asynchronous job which has two endpoints, first to create the job and second to check the status. There is a URL parameter to support creating multiple samples per row. The request body allows for specifying specific rowIds to create samples from, defaulting to all rows. You may also optionally specify sample properties for all samples created as part of the request body.
The following capabilities are in beta and are available for users, administrators and developers on the Signals platform upon request. Please contact your account representative or our support team if you would like access to the following features.
Fundamentals (beta)
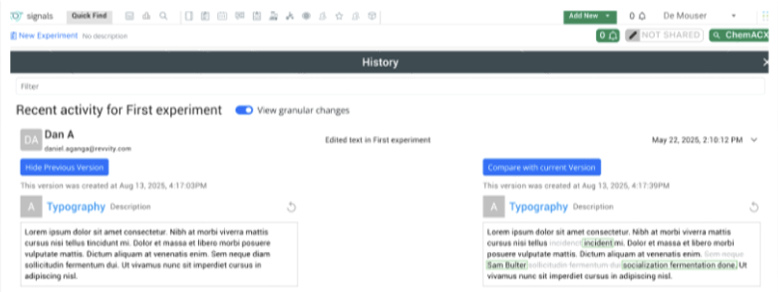
Audit trail displays highlighted granular changes within records for text elements in experiments and experiment-like objects. When users compare an audit record with a previous version, there is a "view granular changes" toggle. Selecting this toggle will show users a clear visual summary: gray highlighting shows text that was edited or removed and green highlighting shows newly added or updated text.
AI (beta)
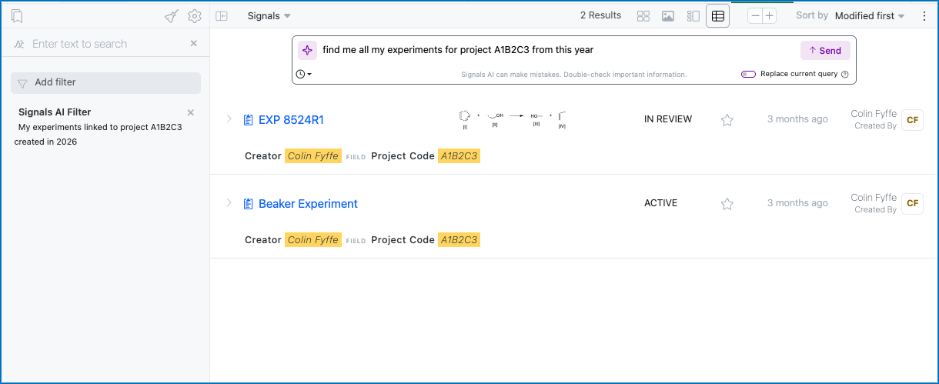
We have added a new Beta AI feature allowing a user in Advanced Search to create a filter using a natural language prompt. This filter will be added on top of your existing filters unless the toggle to replace your existing query is enabled. This is limited to Signals One customers.
Synergy (beta)
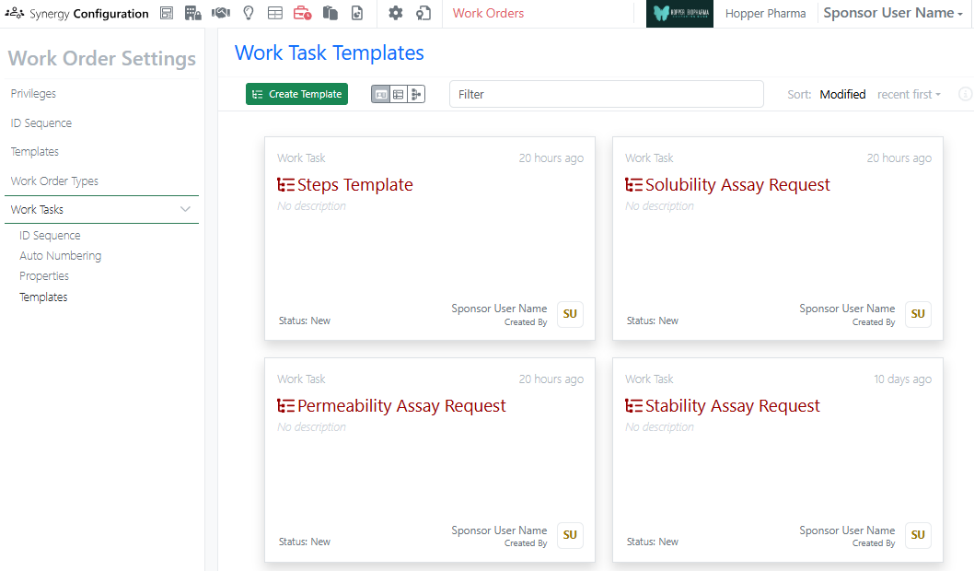
Works Tasks can now be configured and added to Work Orders to create a series of trackable, assignable, and templateable sub-tasks for a Work Order. Work Task properties and templates can be configured in Synergy Configuration > Work Orders > Work Tasks.
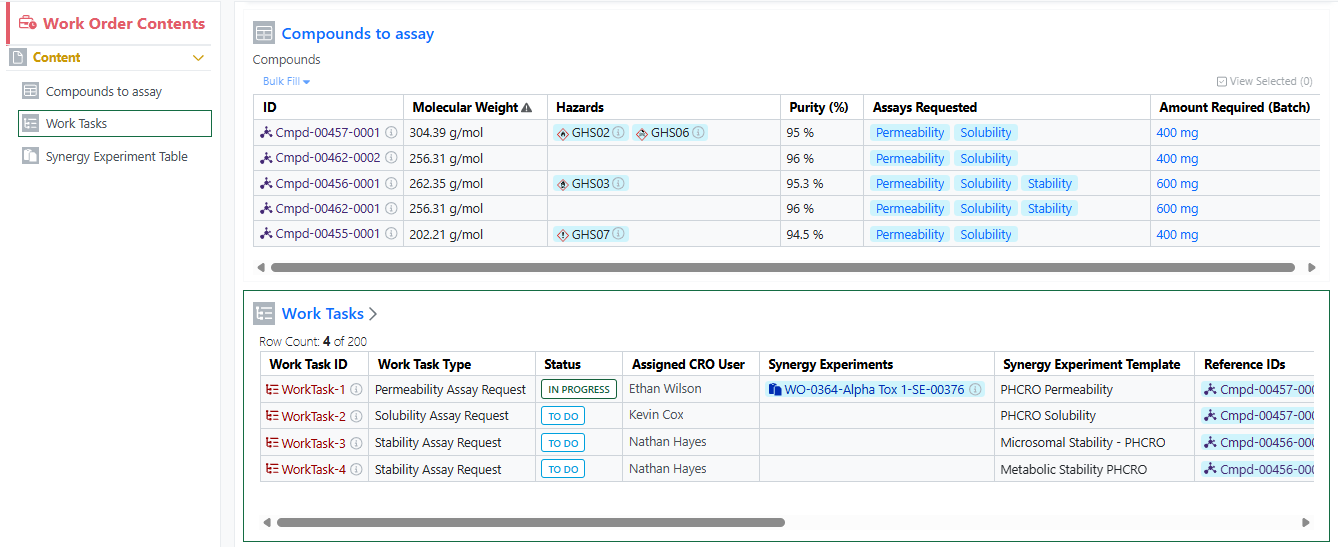
A Sponsor user can add a Work Task to a Work Order when the Work Order is in an editable status (Draft, To Do, On Hold). The Sponsor can optionally specify a Synergy Experiment Template to be used for the Work Task, as well as specify which References the Work Task applies to.
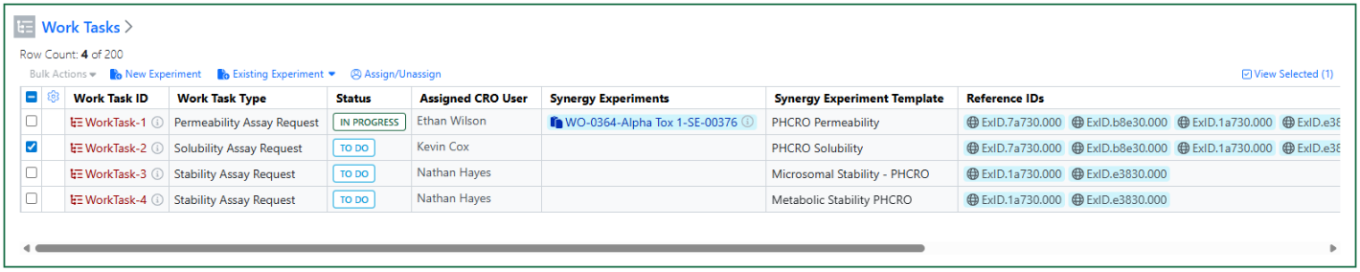
Once a Work Order is sent, Work Task(s) Status changes from New to To Do. A CRO Department manager can optionally assign the Work Task(s) to a CRO user. A CRO user can start one or more Work Tasks by selecting the Work Task(s) and creating a New Experiment or adding the Work Task to an Existing Experiment. The Work Task(s) Status then changes from To Do to In Progress.
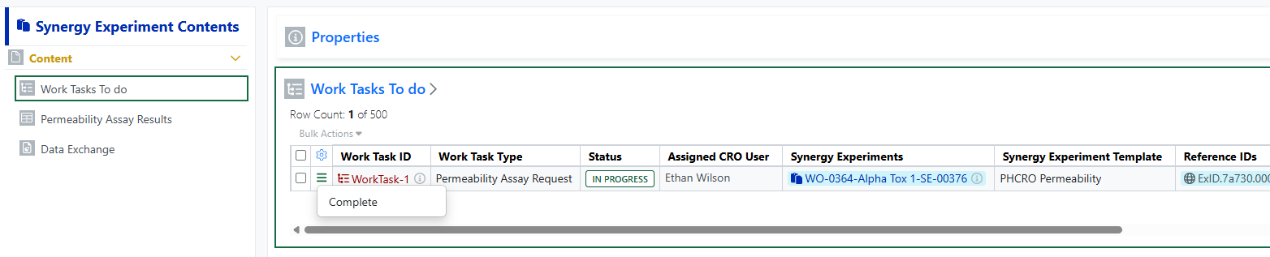
Note, when creating a New Experiment, the specified Synergy Experiment Template will be used when applicable. Also, each Work Task can have multiple Synergy Experiments associated, and additional Synergy Experiments can be added to an In Progress Work Task from the Work Task table in the Work Order.
Work Tasks display in a Work Tasks To Do table in each associated Synergy Experiment. When the task is done, the CRO user can Complete the Work Task. Work Task(s) must be Completed before submitting the Synergy Experiment.
Administration (beta)
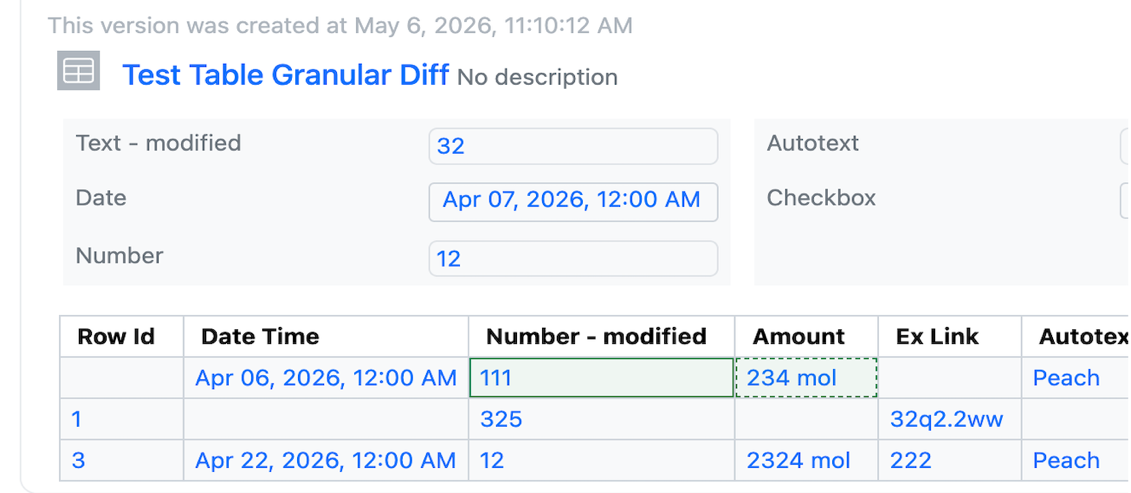
Audit trail displays highlighted granular changes within audit records for admin-defined tables. When admins or users compare a table audit record with a previous version, there is a "View granular changes" option. Selecting this toggle will show users a clear visual summary: gray highlighting shows data that was edited or removed and green highlighting surrounding table cells reflects newly added data. Green highlighting with a dashed border reflects text that was updated.
What's New
We are delighted to release our new version of Signals, 26.3. This release brings enhancements in In Vitro Analysis, Synergy, Inventory, and Chemistry amongst others. We have extended beta capabilities around graphing in tables and curve fit definitions. The beta capabilities around calculated properties for protein complexes are now available to all systems. Finally, we have also fixed a number of small bugs.
The following improvements are available for users, administrators and developers on the Signals platform. Certain features may only be available with appropriate licensing and/or with enablement by an administrator.
- Fundamentals
– The PDF export of experiments can be converted to local time in ISO 8601 format - Data Workflows & Analytics*
– Personal Templates support in the In Vitro Analysis
– Initial support for custom curve fit definitions and publication (beta) - Synergy†
– Move Designs Table to different Idea
– Move Design(s) to different Designs Table or Idea
– Chemical Search for shared Chemical/Design Structures in Reference Tables for Sponsor and CRO
– Reference Table property search for Sponsor and CRO - Chemistry
– Support for pairing of multiple RNA/DNA strands to a single sequence
– Structure searching for monomers in HELM Monomer Curation application - Tables
– End user adjustment of Scatter Plots (beta) *
– Export of image from Scatter Plots (beta) *
– Highlight selected rows or points in the Scatter Plot and Table (beta) *
– Conditional calculations can be contained within conditional calculations
– Increase component limit from 30 to 50 in the Variation Table - Samples
– Empty Sample tables in Templates - Inventory
– Allow Multiple Additions of the Same Container in Materials Table and Stoichiometry Table
– Add multiple (same) sublocations to a parent location easily
– New Labels - InventoryGo ‡
– Reconciliation and Rectification
– Move
– Consume - Materials
– Calculated properties for protein complexes available to all systems
– Inherited Properties for materials available to all systems
– Log P, predicted pKa, and additional compound calculated properties
– Configurable display units for calculated molecular weights - Search
– New options for circular and linear Biological sequence searches - Administration
– The 'created at' field in the system audit trail CSV export can be converted to local time in ISO 8601 format
– Spotfire for Signals Custom Apps Configuration available for Private Cloud exports Spotfire for Signals Online Debug Logs * - Integrations & APIs
– Selected rows in an Hierarchical Table can be identified in an External Action *
– External Action for Edit Reviewers
– API to create a Reference Table in a Work Order
– API to add a reference(s) to a Reference Table
– API for date based data extraction for SDF
– Additional Chemical Structure formats for updating Materials
– PKCE Support for Bearer Token Exchange
* Requires Signals One license
† Requires Signals Synergy license
‡ Requires InventoryGo license
We also fixed several small bugs in this release. Details of the enhancements are described below.
Administrators should be aware that we have retired most of the In Vivo Apps from the Signals One App Store in Spotfire. Specifically the following Apps are no longer available: Study Designer, Dose Preparation, Baseline Capture, Sequence Of Events, Sequence File Generator, Mass Spec, Dose Verification, Business Rules, and Data Model Designer. The PK Parameters App continues to be supported and will remain in the Signals App in Spotfire in Signals One.
Administrators should be aware that we will have retired the CRAIS Checker capability from Signals Notebook and Signals One. This capability, which was only available via request, was built to allow integration specifically to the Patcore CRAIS Checker database. Such workflows have been superseded by the Chemical "External Checking" capability which can be used against any chemical compliance database.
Data Factory Administrators should note that legacy Data Factory projects will be fully deprecated in a future release. A straightforward migration path will be provided in an upcoming release to support this transition.
Data Factory administrators should also note that the Advanced Security policies in Inventa apply an "AND" logic across all user groups, restricting access if any groups lack permissions. This behavior differs from the standard Signals security model and will be fixed to align with the standard in a future release.
Administrators are recommended to subscribe to the channels within our support news site found at https://support.revvitysignals.com/hc/en-us/categories/360004446171-Support-News which contains more information about releases and other pertinent product information.
This content is anticipated for release on our Production E3 environments, and for Private Cloud customers on our deferred release schedule, in July 2026.
Further Details
The following improvements are available for users, administrators and developers on the Signals platform. Certain features may only be available with appropriate licensing and/or with enablement by an administrator.
Fundamentals
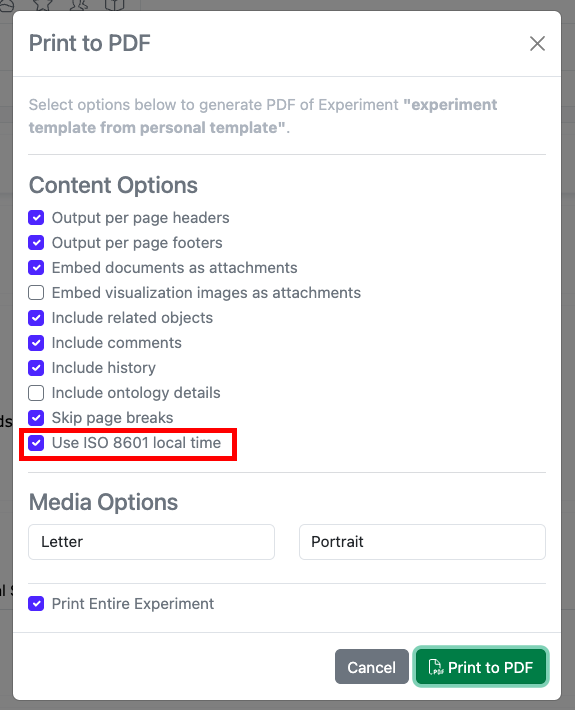
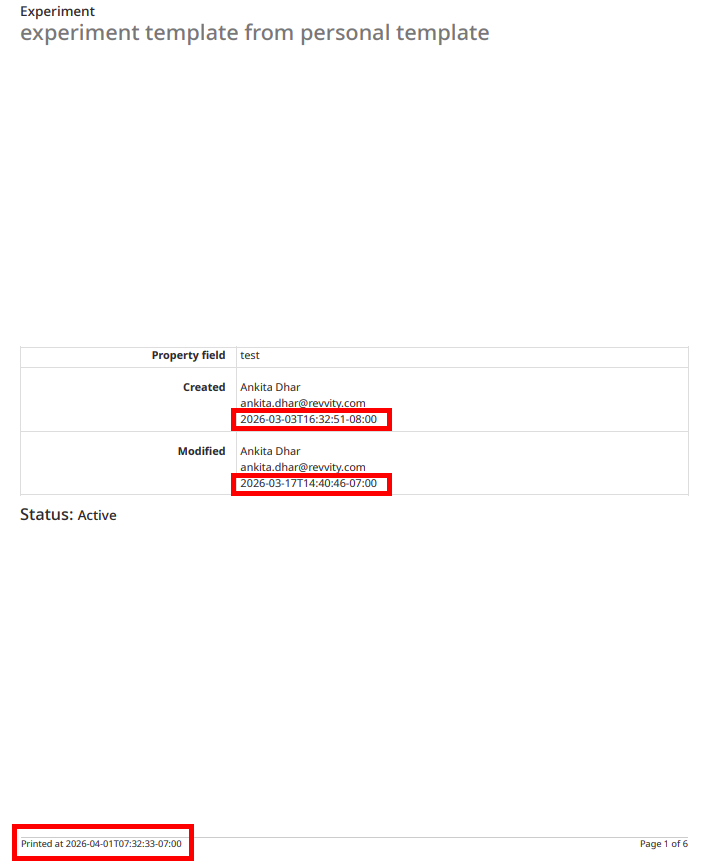
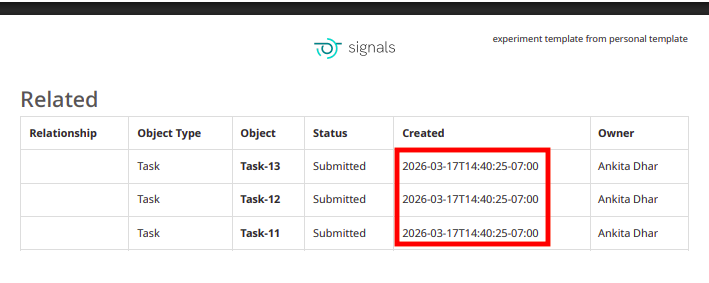
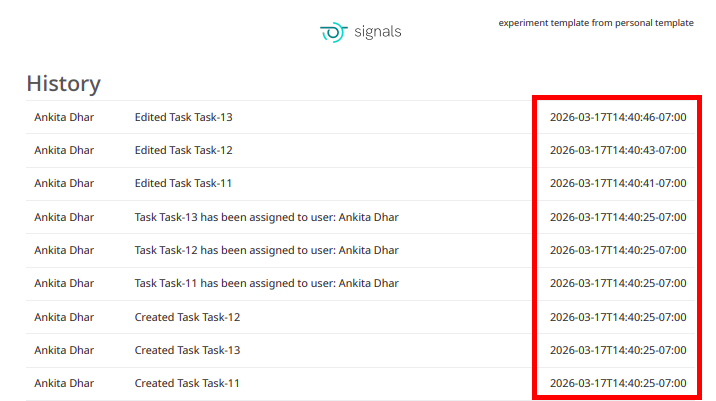
The PDF export of experiments can be converted to local time in ISO 8601 format. When end users select the ‘Print to PDF’ option in an experiment, a new setting can be selected to “Use ISO 8601 local time” which will convert timestamps throughout the PDF to local military time with a UTC offset.
Data Workflows & Analytics
In Signals One, In Vitro Analysis users can now save an instance of an In Vitro Analysis object as a personal Template:
Saved In Vitro Analysis personal Templates become then available for the user for subsequent new analysis.
Synergy
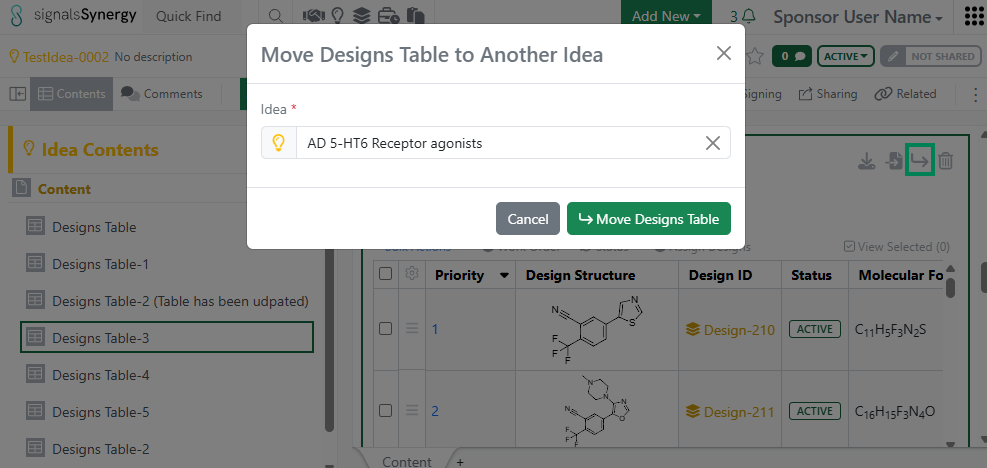
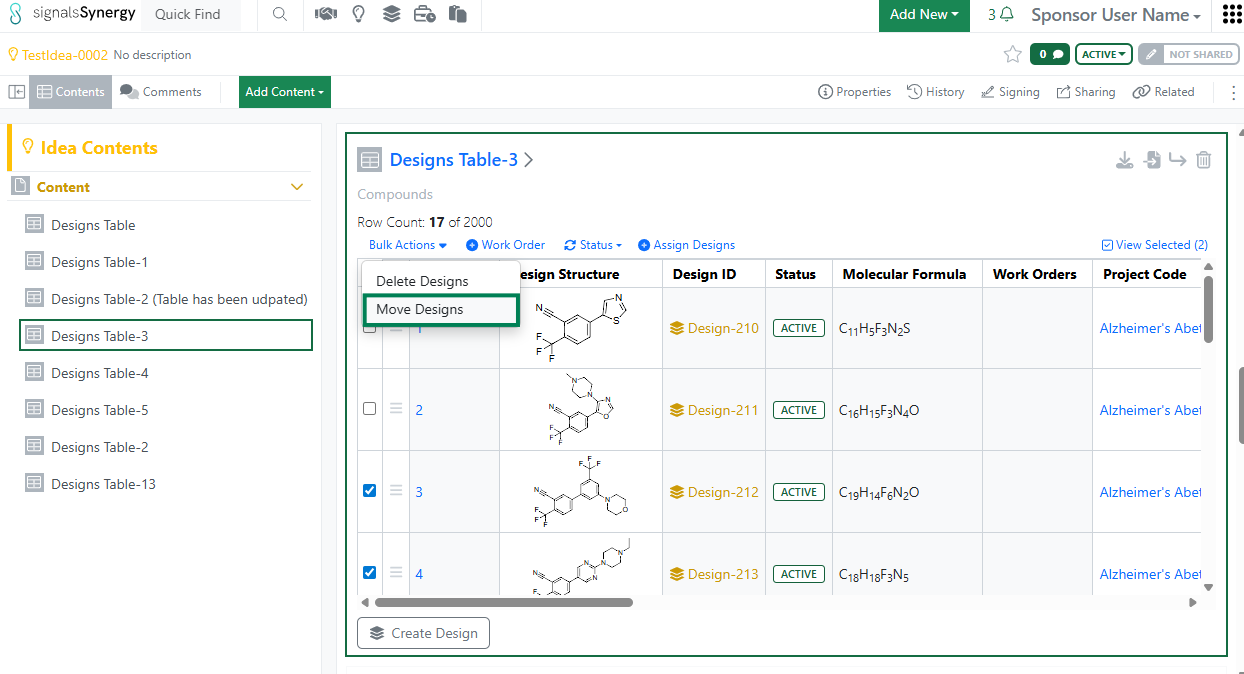
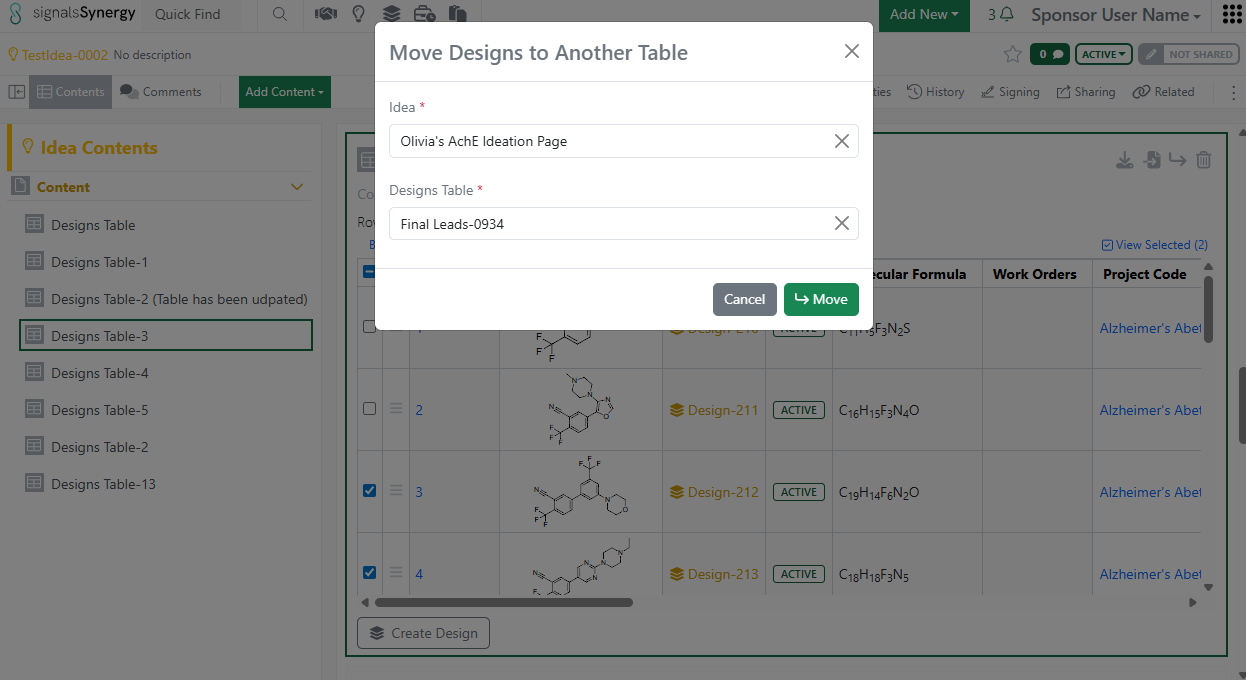
Given the new privilege to Move Designs, a Sponsor user can now move a Designs Table to a different Idea or move selected Design(s) to a different Designs Table. The user must have Full Control to an Idea in order to move Designs.
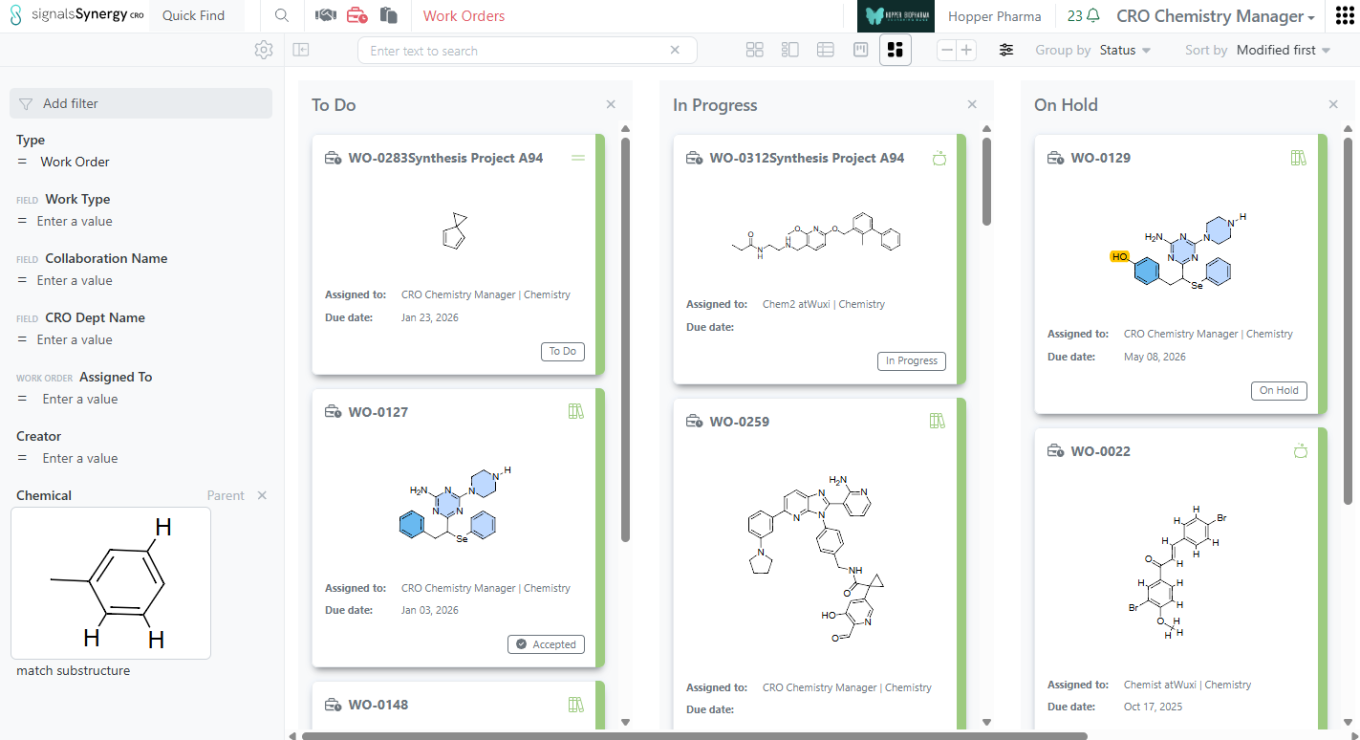
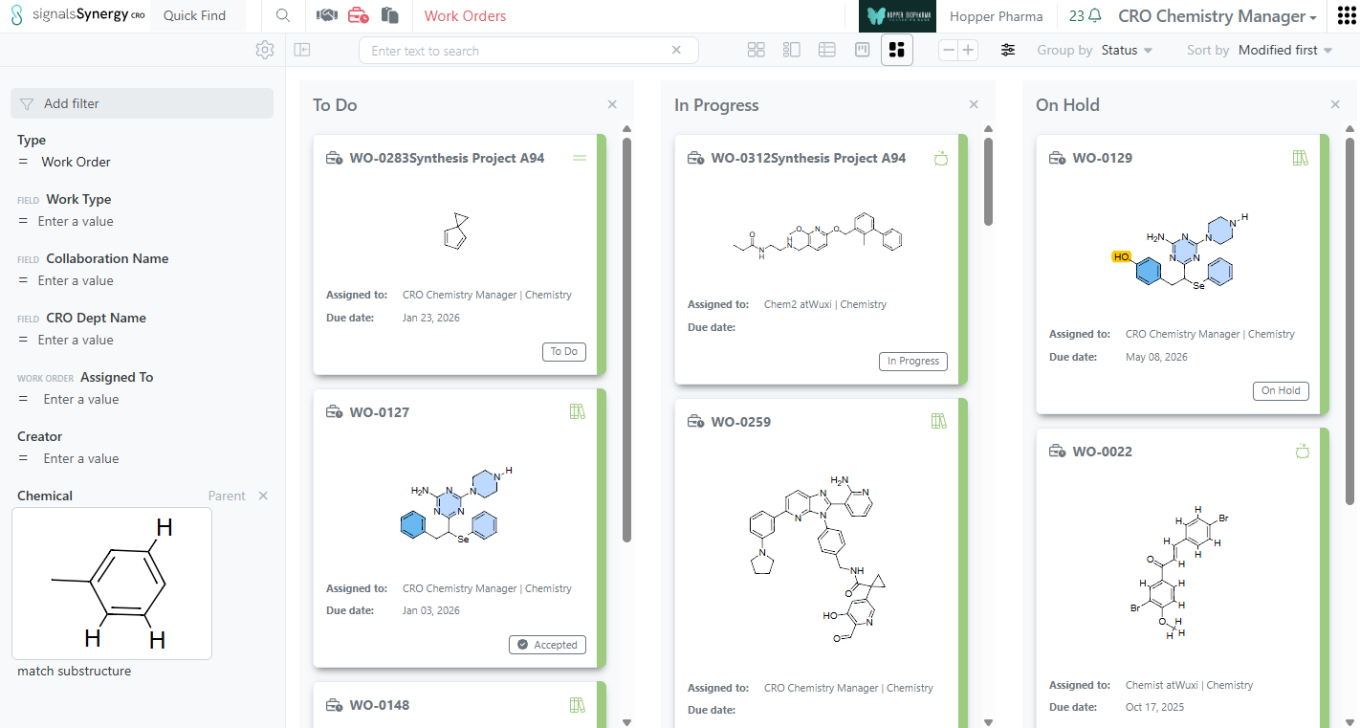
As a Sponsor or CRO user, I can now search for my Work Orders based on the content of the Reference Table (e.g., list of Designs to be made or batches to be tested). Reference Tables allows the Sponsor to selectively share a sub-set of Design or Material Asset or Batch properties when requesting Design synthesis or assays to be run. If a Design Structure or Chemical Structure is included in the properties shared to the CRO, the Sponsor and CRO can use Chemical search to find the Work Orders containing a specific structure or sub-structure within Advanced Search or the Work Order Smart Folder. Note that the Chemical filter should be set to Parent to ensure that the Work Order containing the Reference Table is found.
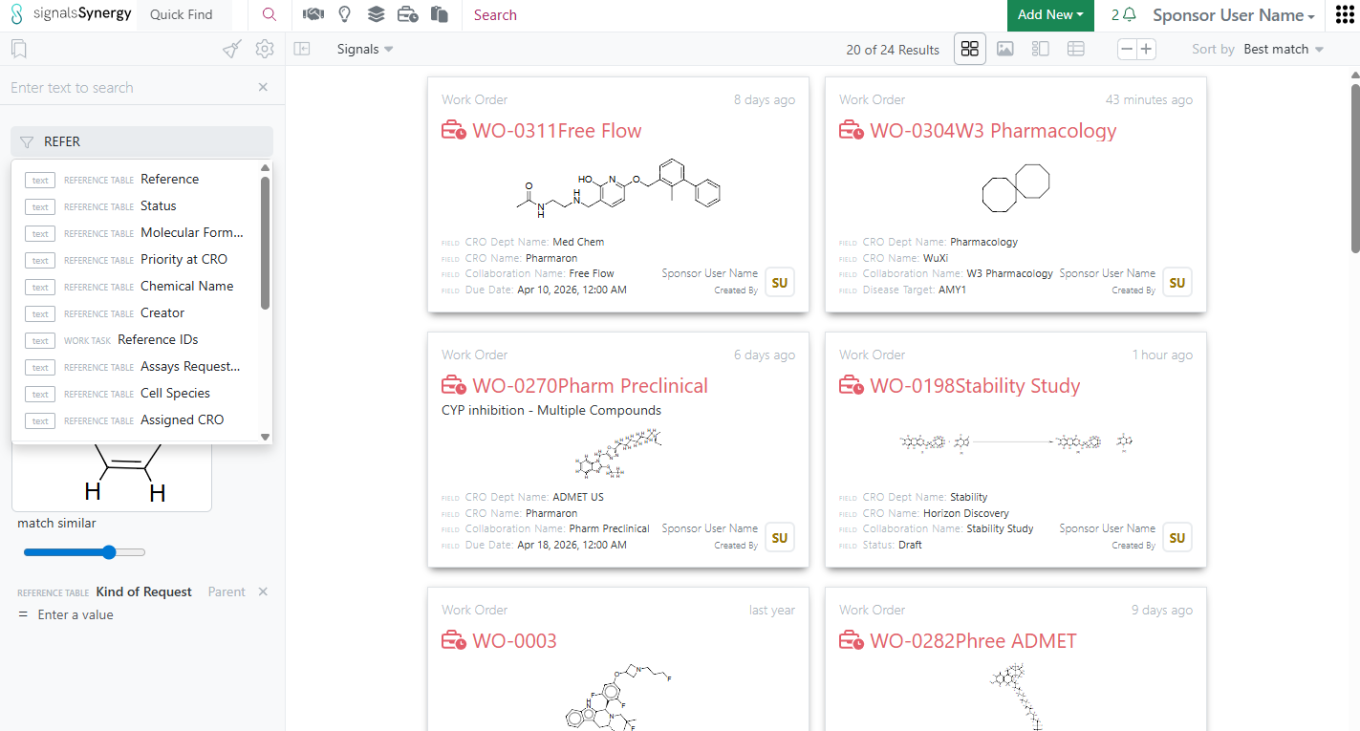
In addition, non-structure properties shared in the Reference Table can also be used when searching for Work Orders within Advanced Search or the Work Order Smart Folder.
Chemistry
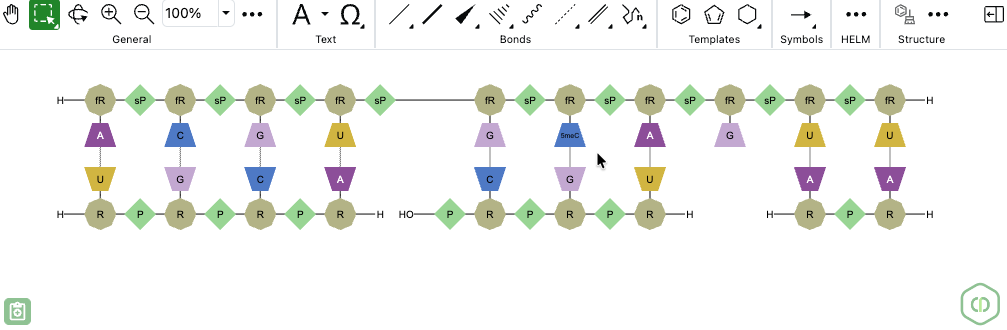
ChemDraw now allows users to draw biopolymers with multiple complementary RNA strands for a single sense strand. When a set of hydrogen-bonded RNA sequences is created, ChemDraw detects the sense strand, identifies each first-degree complementary strand, and automatically places each complement beneath the sense strand according to hydrogen bond positions. The sense backbone is stretched as needed to minimize overlap, enabling side-by-side display of complements with proper alignment.
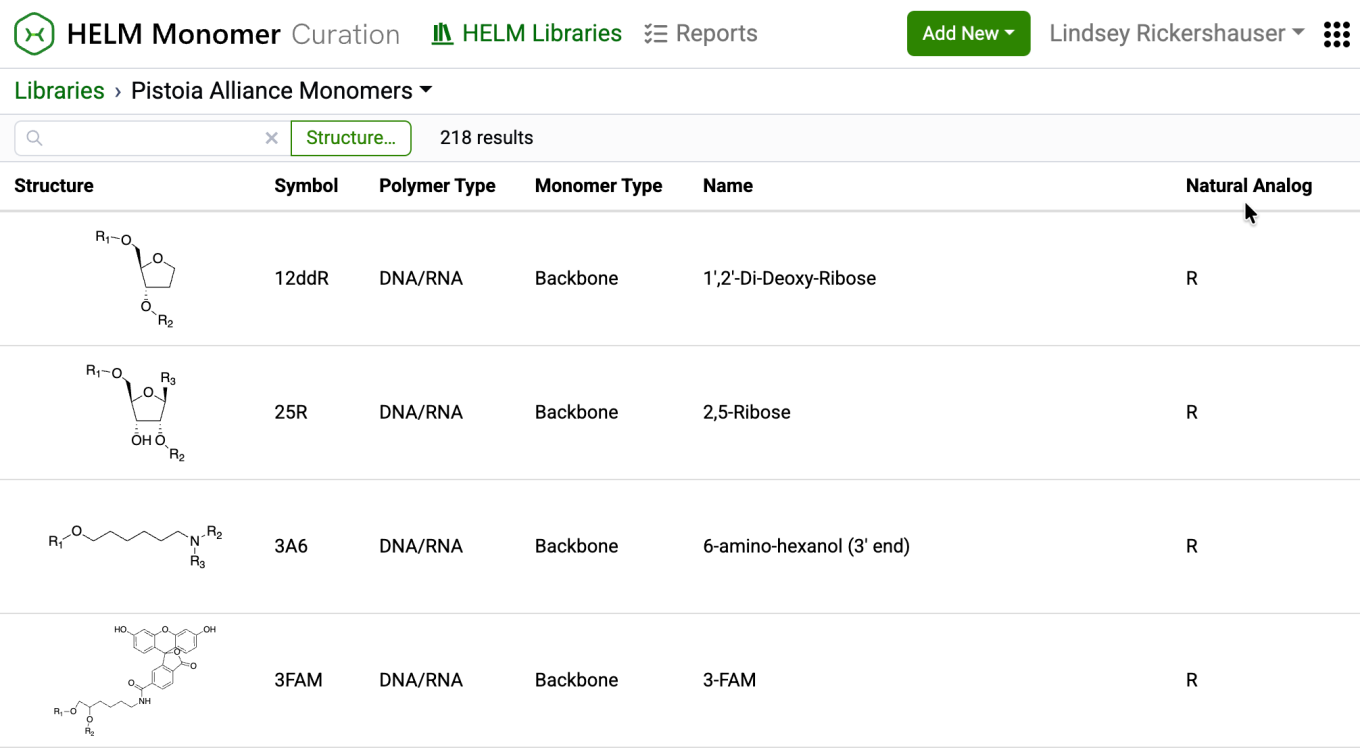
New capabilities have been added to the HELM Monomer Curation application for filtering and searching monomers based on chemical structures. Available as a button next to the text search field in the HELM Libraries list view, the ‘Structure…’ button opens a structure editor.
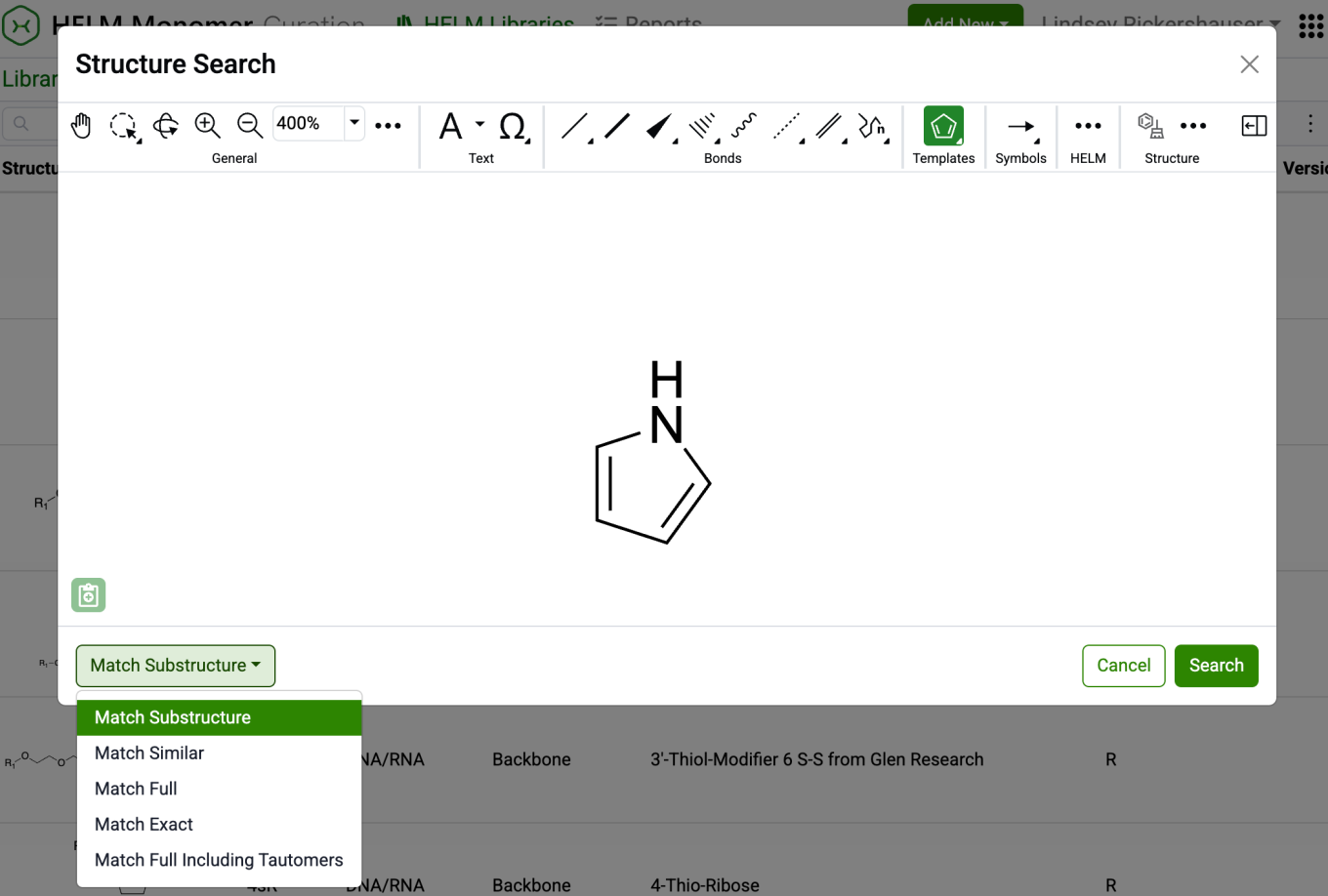
After drawing a structure, users can select from substructure, similarity, full, and exact search options. Selecting ‘Search’ will then initiate the search filter.
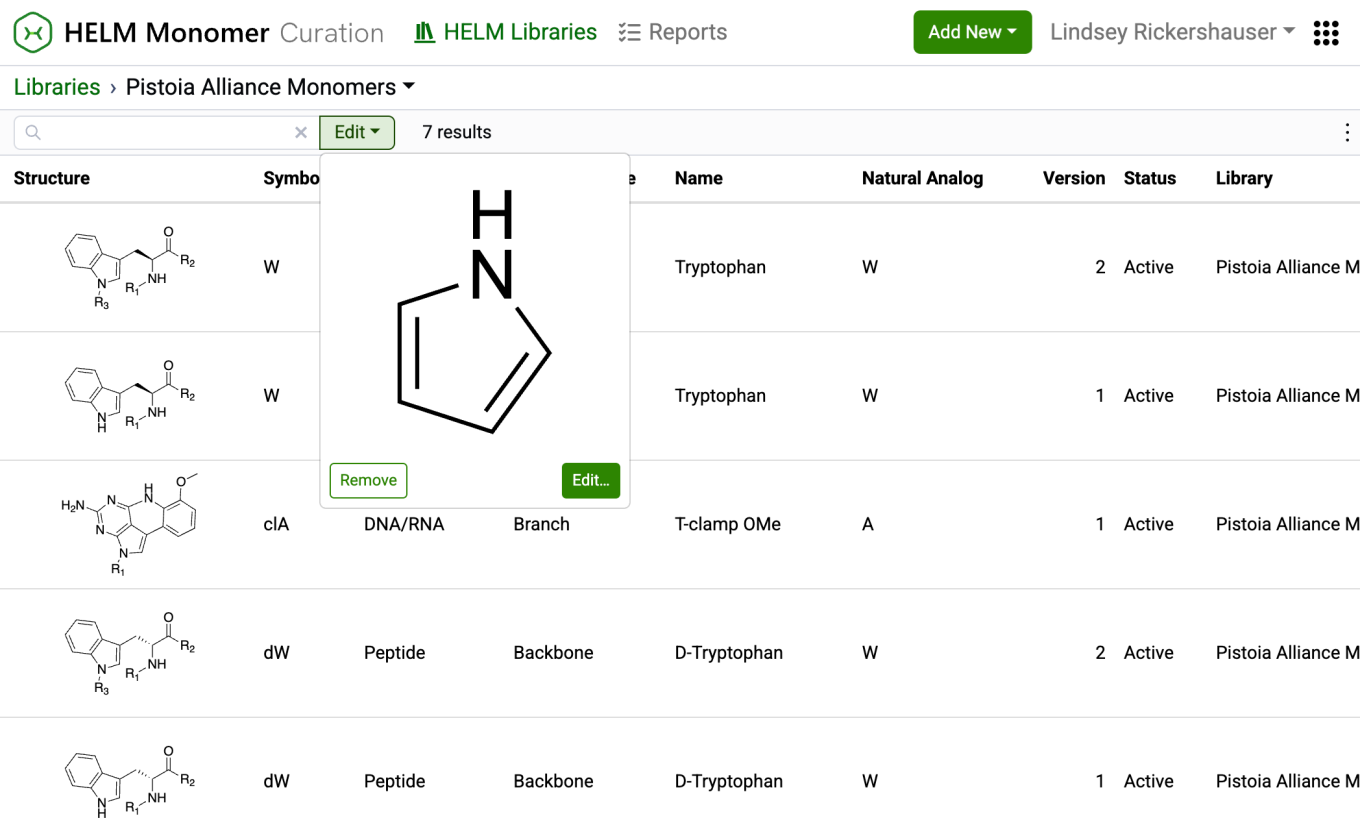
After completing a structure search, the ‘Structure…’ button transforms into an ‘Edit’ button. Selecting this button allows for previewing the structure, making changes to it, or removing it from the search filter. Structure searches can be combined with text-based searches to refine results even further.
Tables

Conditional calculations are now allowed to be used within other conditional calculations, such as within a CASE statement. For example a CASE statement can use another conditional statement such as an IF for each clause. A use case is to enact specific aggregate calculations over restricted values, such as if the sum of a given component in a formulation, such as water, is needed for an additional calculation.
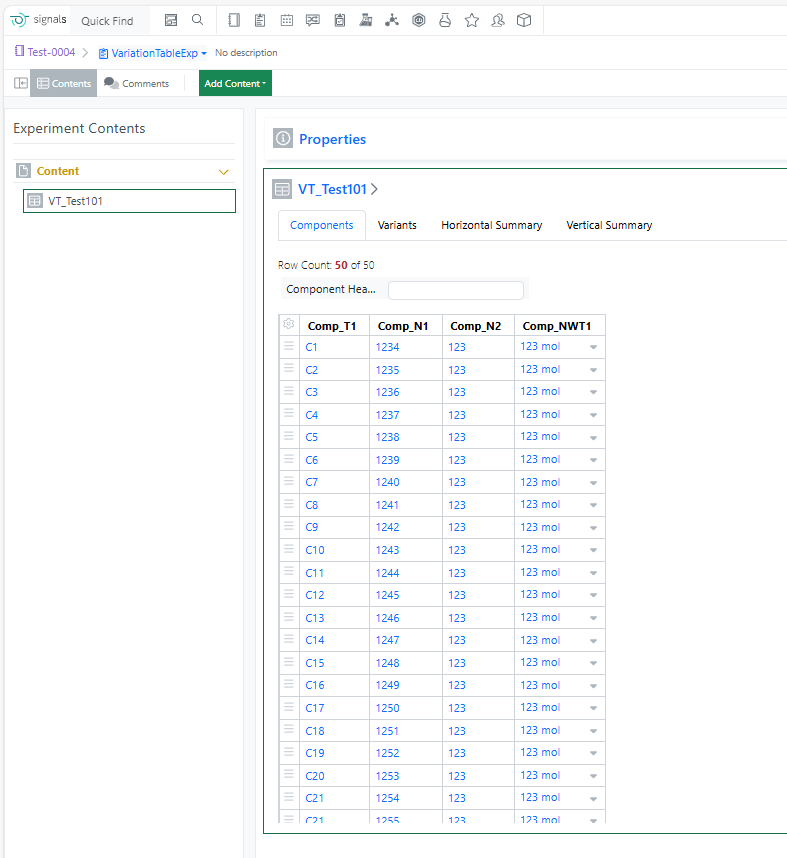
The component limit in the Variation Table has been increased from 30 to 50, enabling more complex experimental designs and reducing the need to split large variations across multiple tables.
Samples
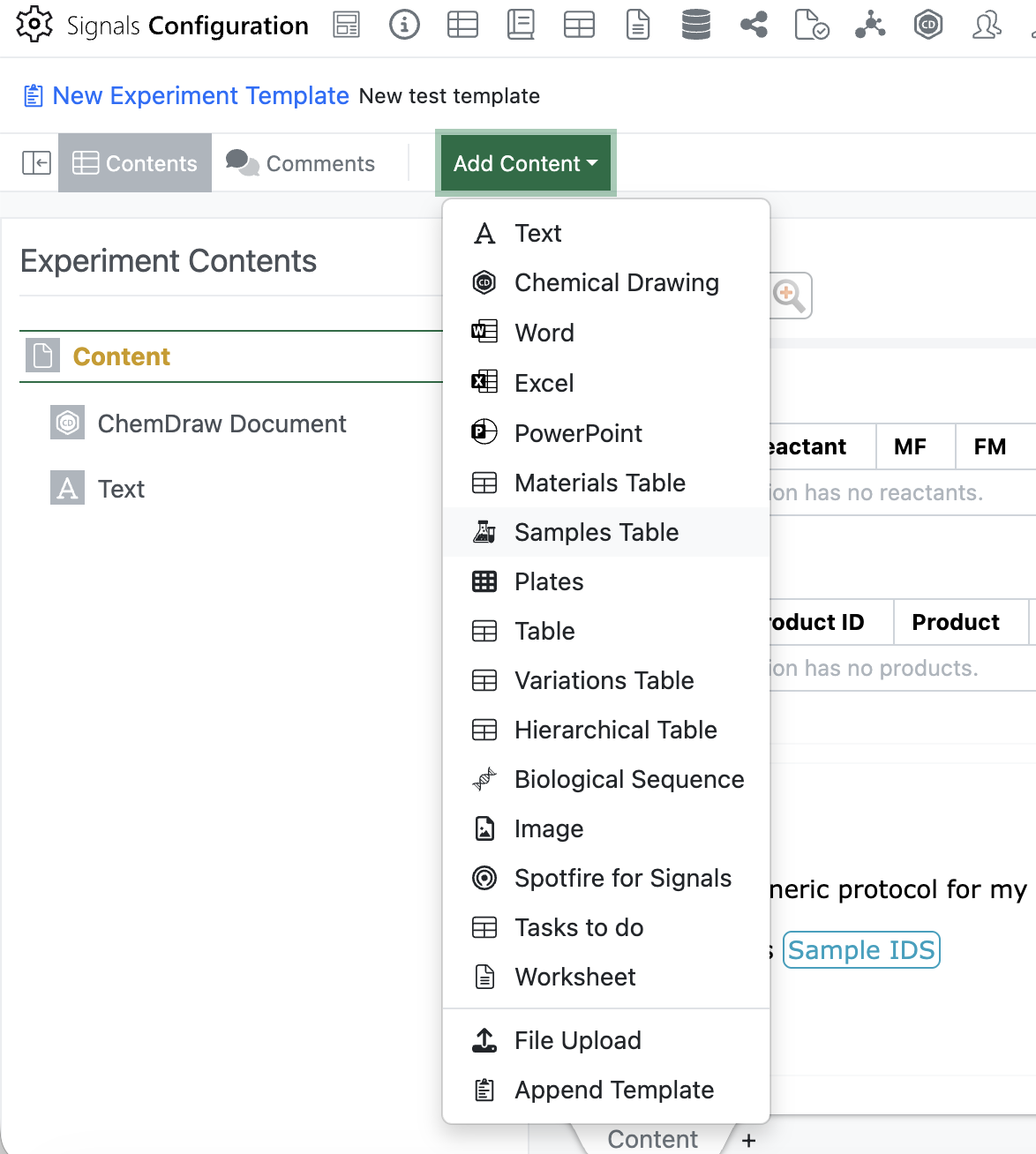
Admins will now be able to add an empty Sample Table to an Experiment or Experiment-like object template using the “Add Content” menu. Only one sample table is allowed in each template, once the sample table has been inserted it will be removed from the “Add Content” menu.
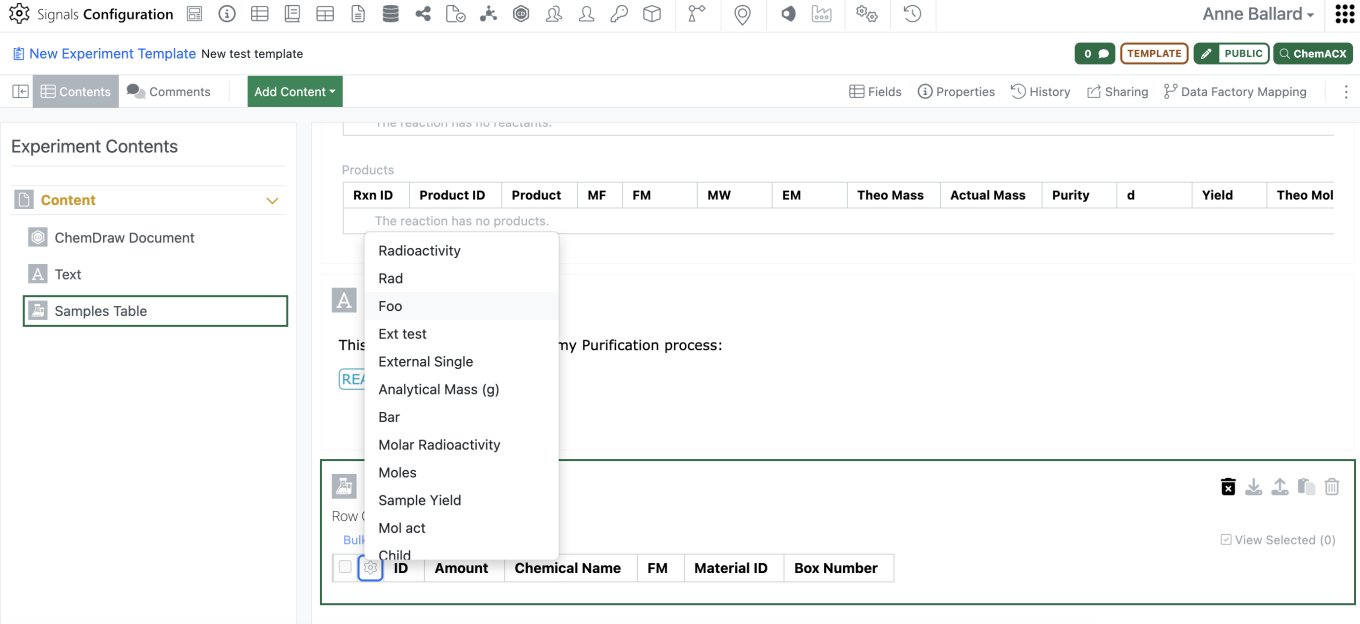
Admins should be able to then move the empty sample table to wherever in the template they would like to have the Sample table. Admins are also able to preselect the columns for the sample table in the template using the gear icon in the left corner.

For an end user, when creating the experiment, if there is an empty sample table in the template the empty sample table will appear in the experiment. When the user adds a sample, it will be added to the existing sample table with the selected columns as defined in the template. This will allow admins to vary which sample columns are shown from template to template without utilizing multiple sample templates.
If a user deletes all samples from the sample table, they will now need to delete the empty sample table with an additional click. The empty sample table can be included in the experiment copy function without including the samples.
Inventory
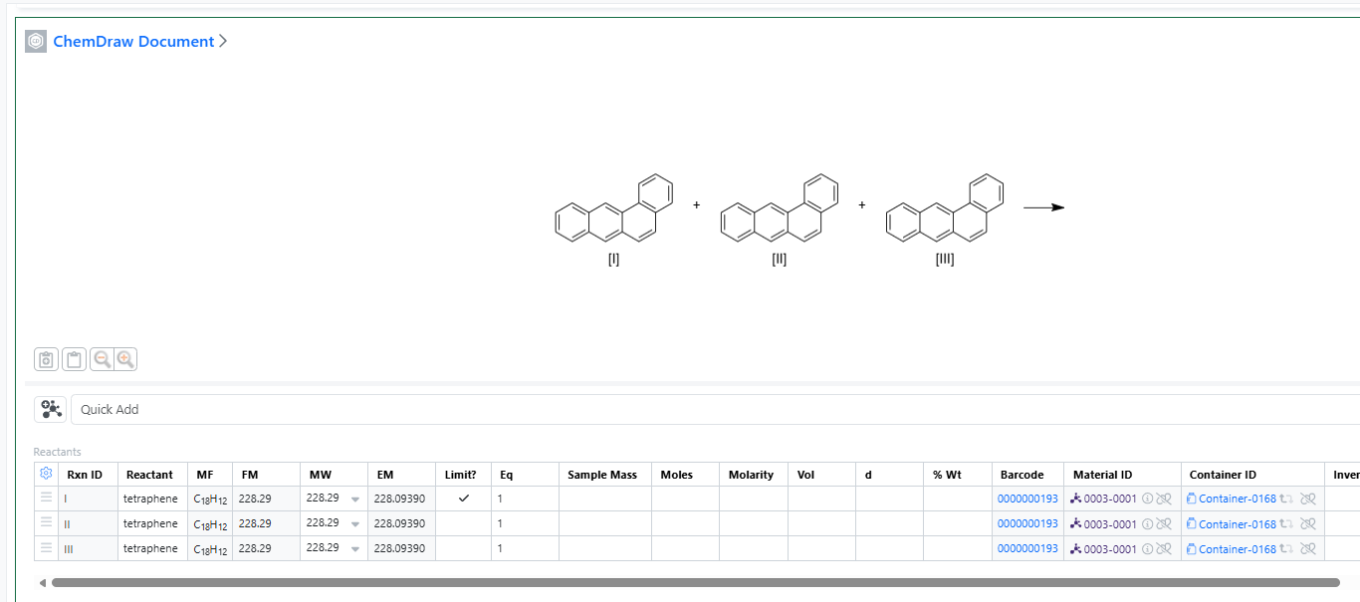
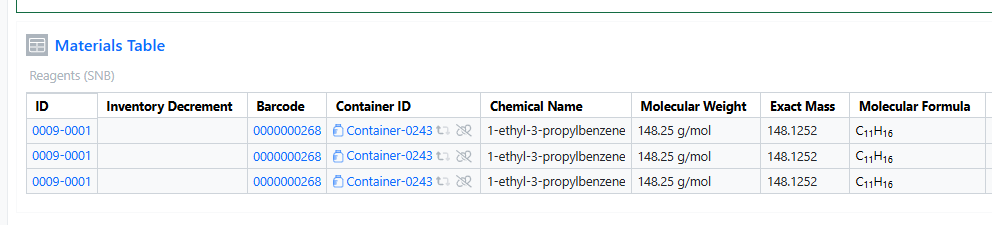
You can now add the same container multiple times in both the Materials and Stoichiometry tables. This enhancement improves procedure documentation clarity, stoichiometric accuracy, and inventory traceability, especially for workflows involving repeated dosing, split additions, or multi‑step reactions.
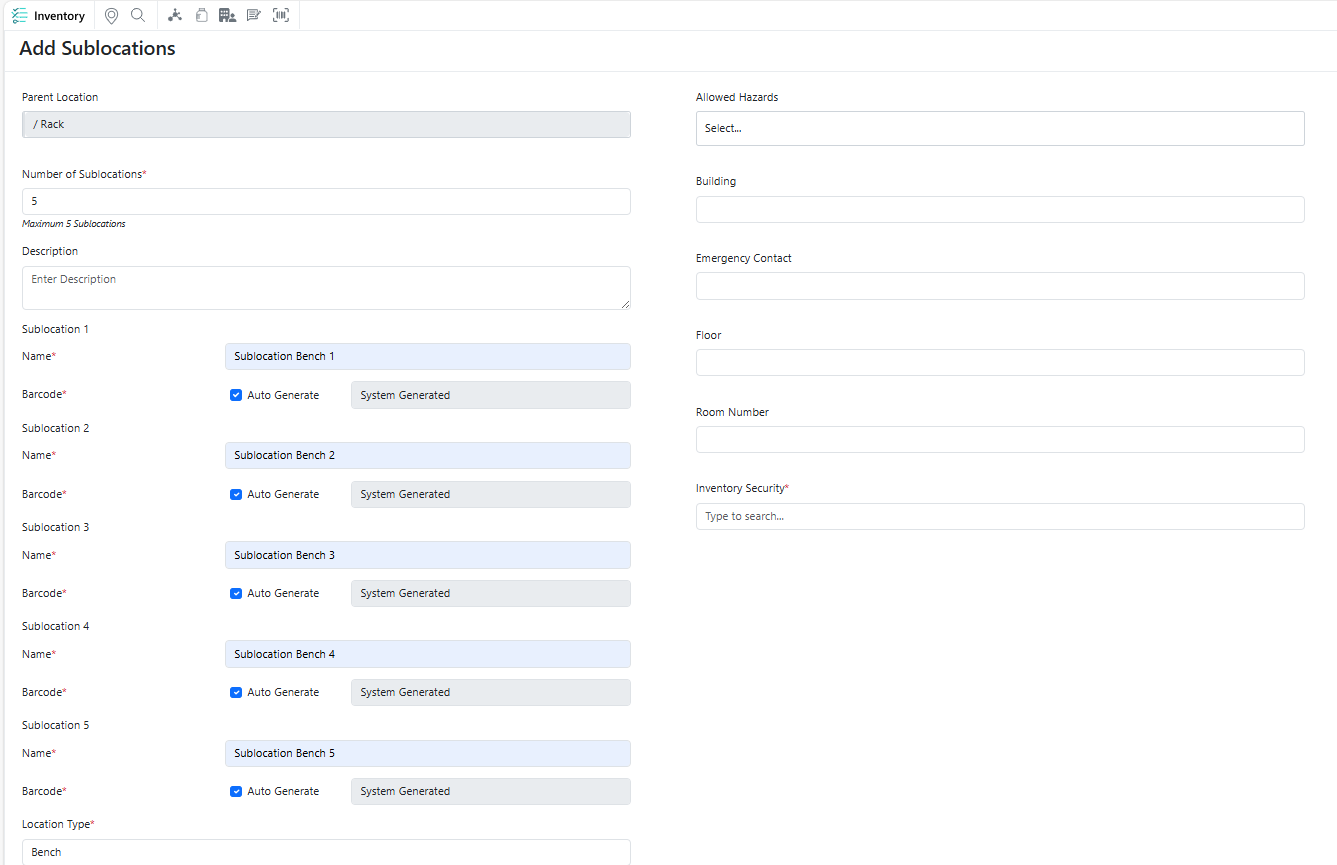
You can now quickly create multiple identical sublocations under a single parent location, making it easier to build and manage hierarchical location structures at scale while reducing repetitive setup effort.
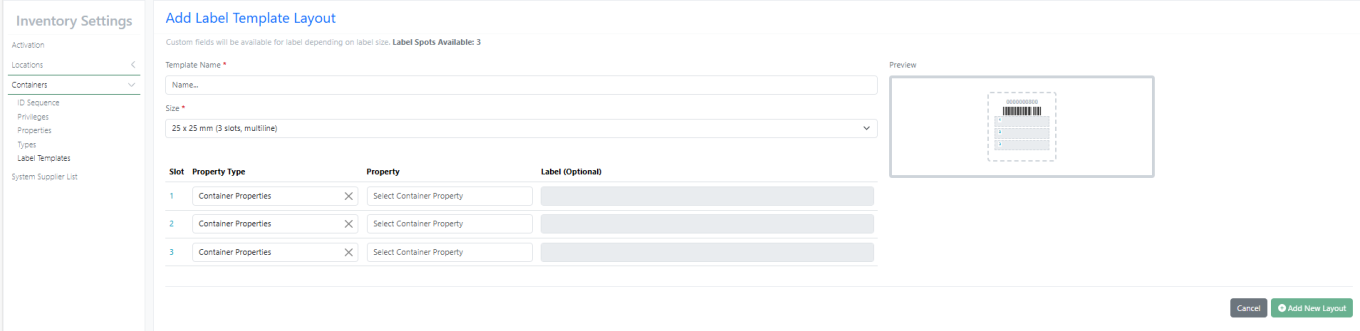
New Label – 25 X 25 mm (3 slots, multiline) supports multiline with font size 6
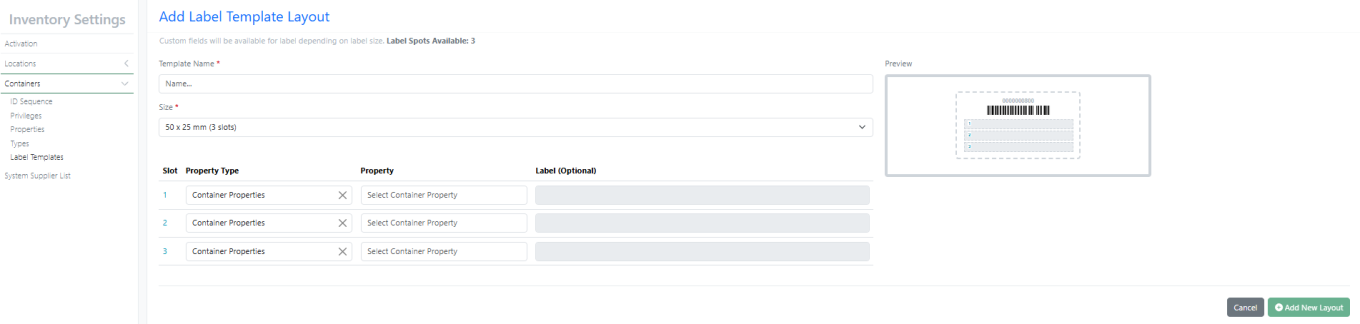
New Label – 50 X 25 mm (3 slots) supports multiline with font size 8
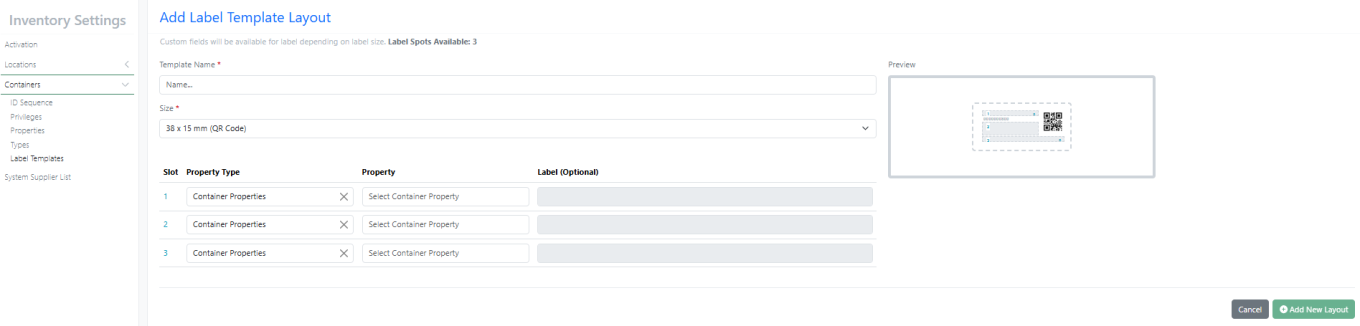
New Label – 38 X 15 mm (QR Code)
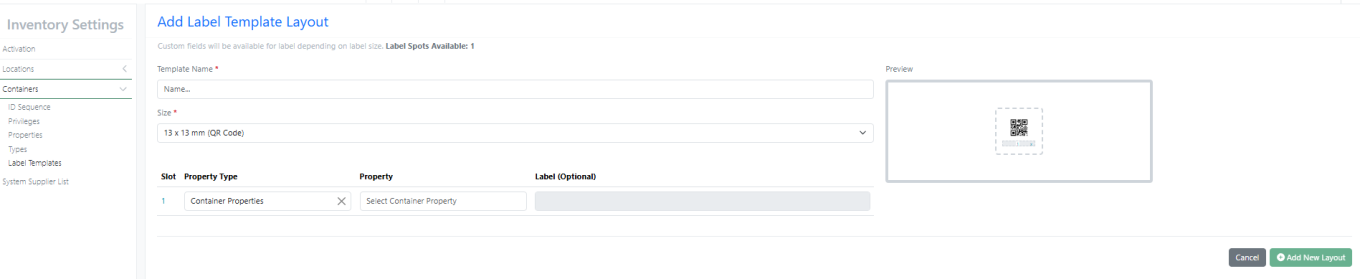
New Label – 13 X 13 mm (QR Code)
InventoryGo

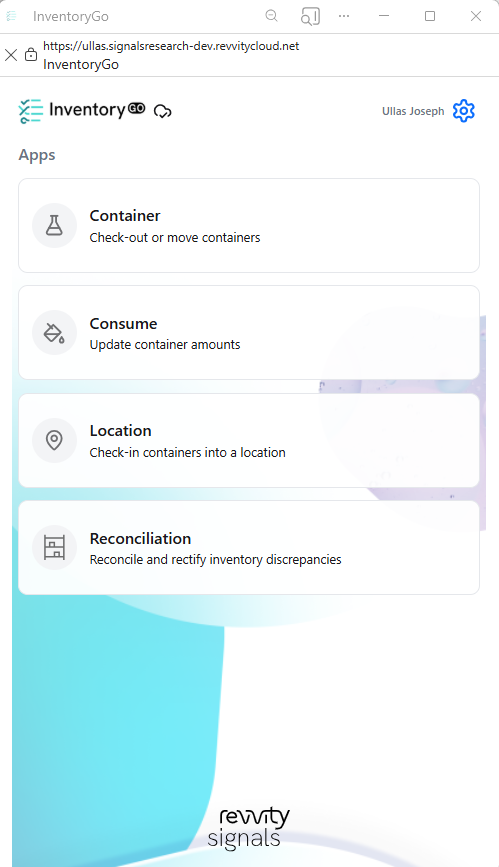
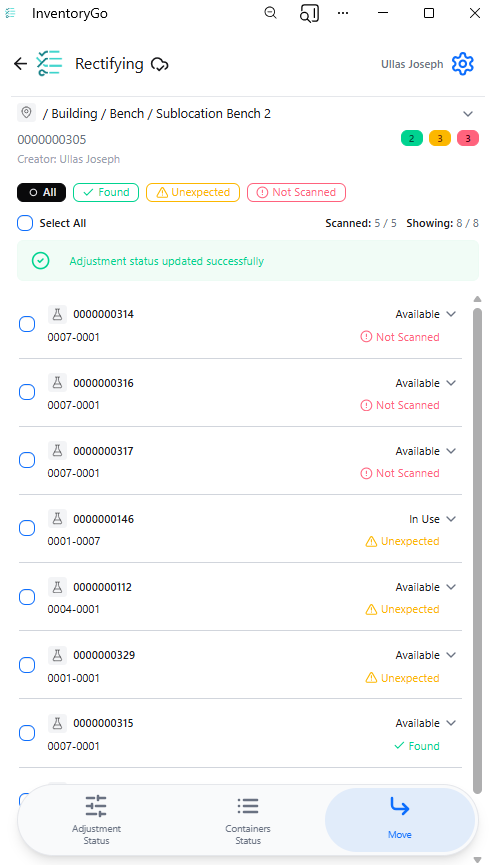
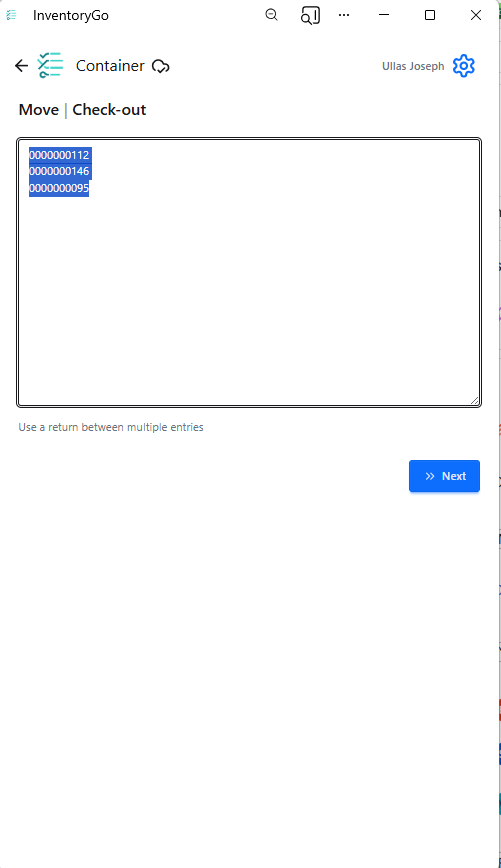
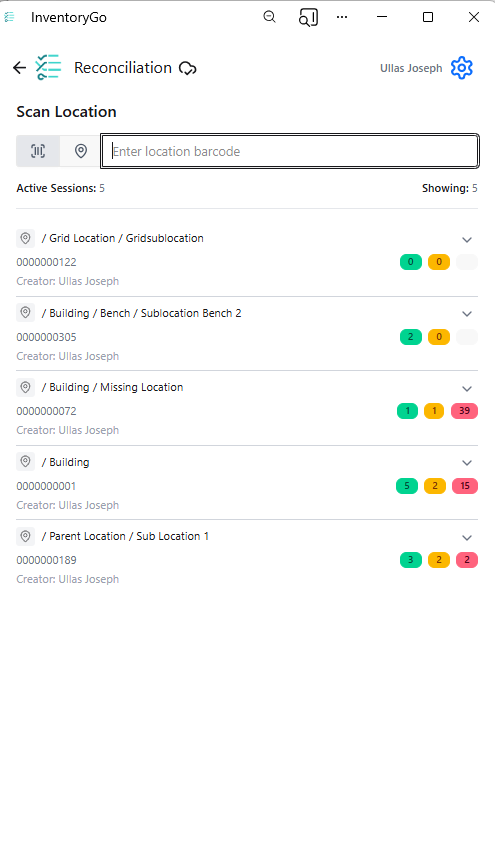
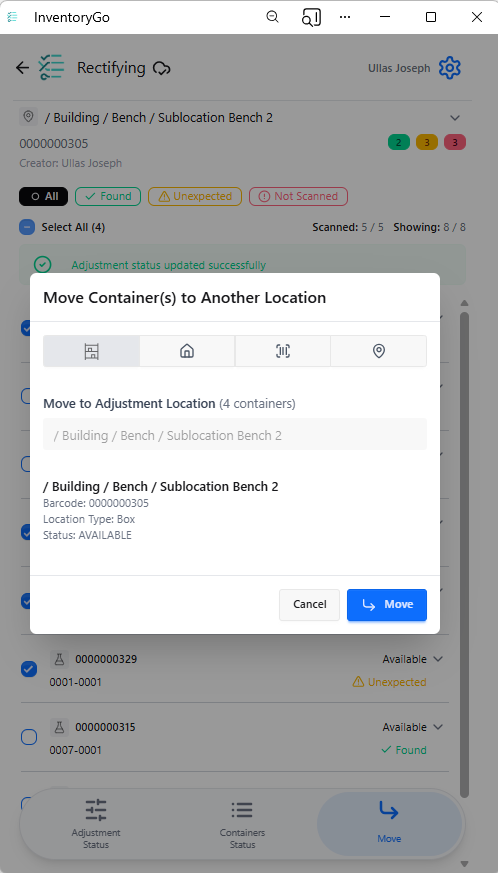
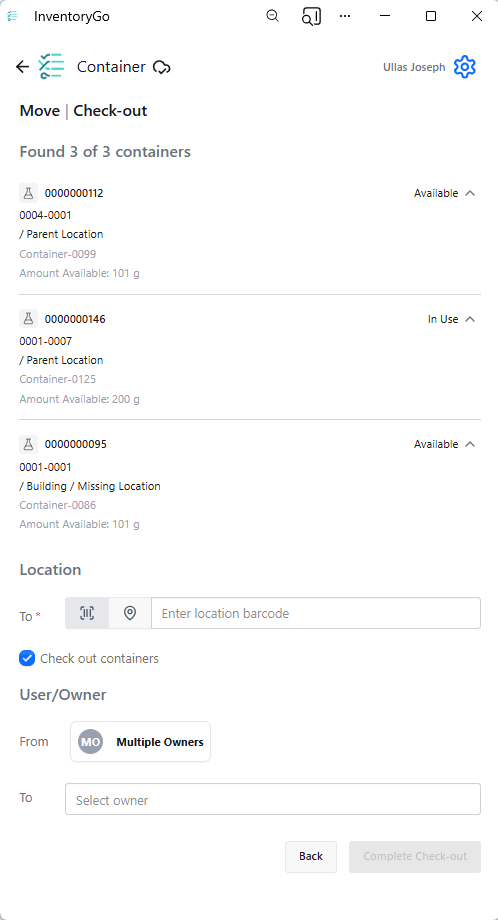
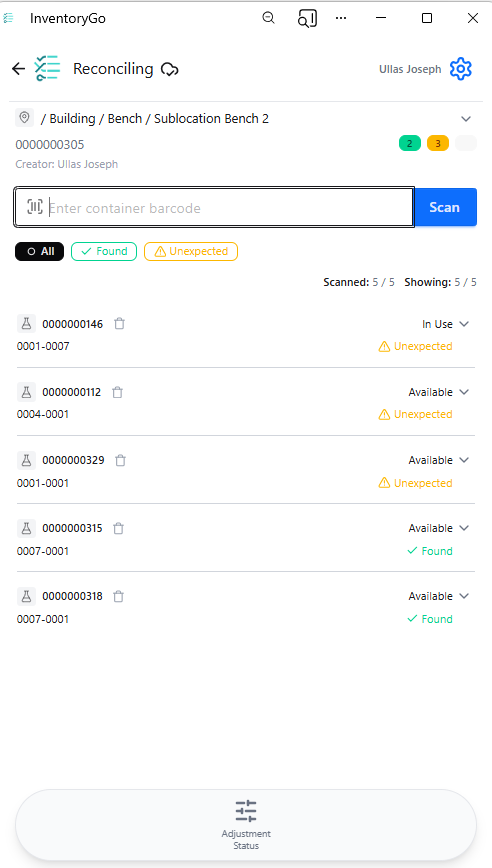
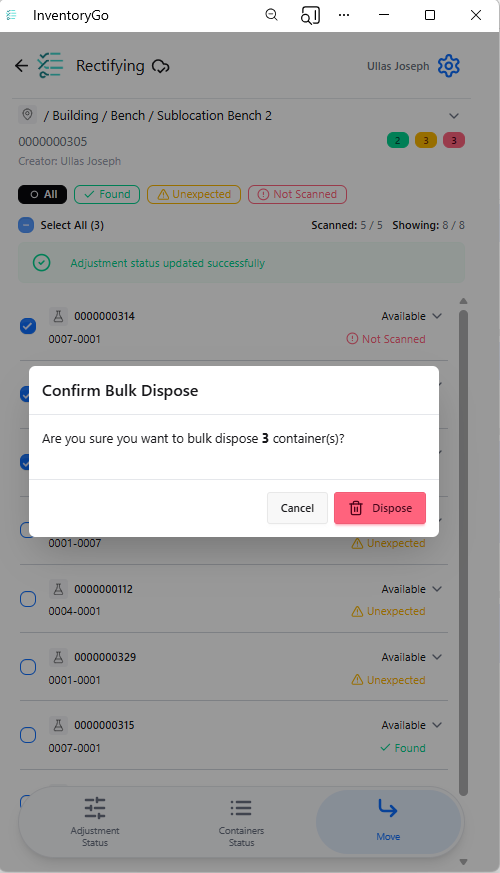
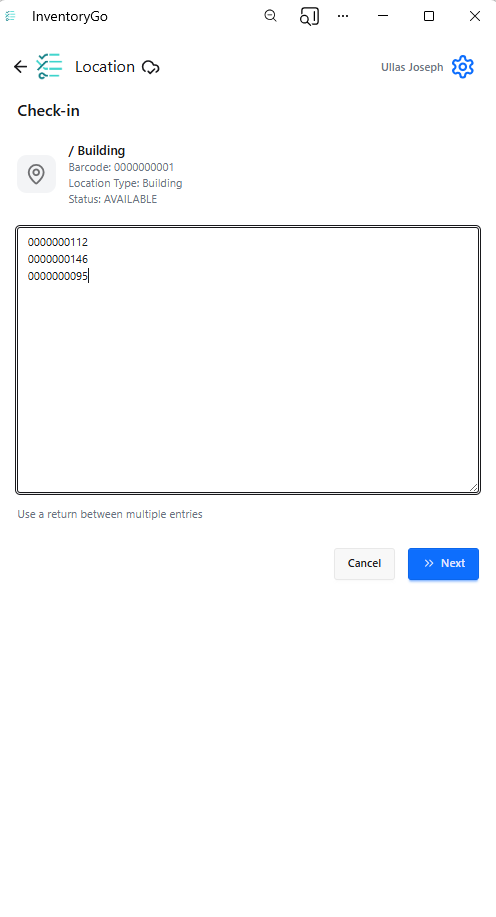
InventoryGo introduces mobile‑first inventory workflows that enable Reconciliation and Rectification, along with Move and Consume actions directly from the lab or storage location. Users can quickly reconcile physical inventory against system records, correct discrepancies, move containers between locations, and update consumed quantities through a streamlined, scan‑based experience—improving inventory accuracy, productivity, and audit readiness. InventoryGo is available as an add‑on capability. Please contact your Sales Associate for more information on enabling InventoryGo for your tenant.
Materials
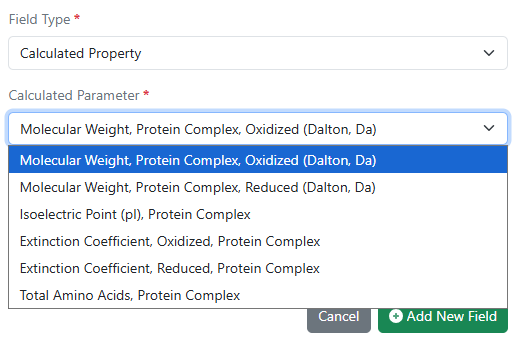
Calculated properties for protein complexes for Custom and Protein material types and the preconfigured Antibodies library are now available to all systems. Molecular weight, isoelectric point, extinction coefficient, and total length can now be computed for complexes comprised of multiple protein parts linked to via internal references.
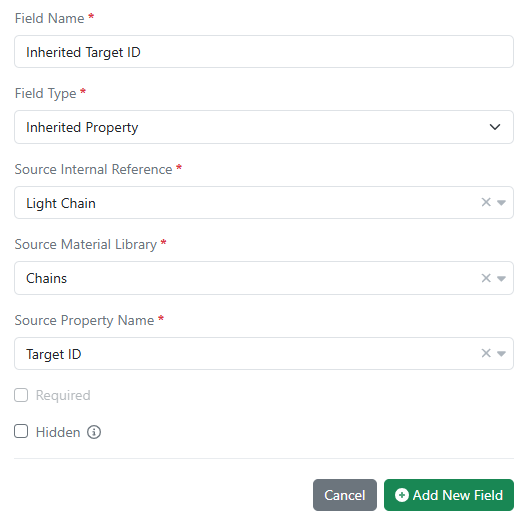
Inherited properties in all material libraries are now available in all systems. Data field values can now be passed down (i.e. inherited) from one material to another using internal reference links. Updates to the source data are automatically propagated to any inheriting entity and are kept in sync.
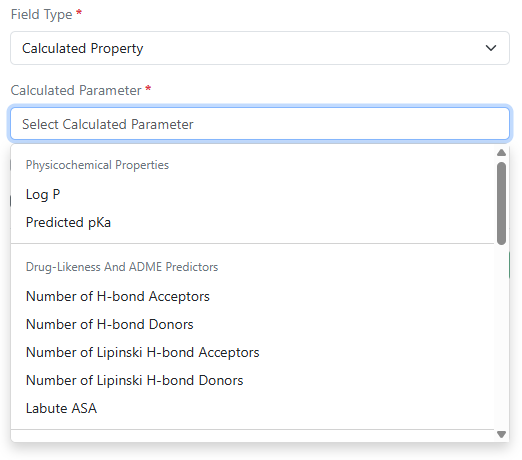
Log P, predicted pKa, and additional calculated properties are now available in Compound and Normalized Compound material types and the preconfigured Reagents (SNB) library. Administrators can selectively choose from a variety of physicochemical properties, drug-likeness and ADME predictors, structure and topology, ring system, atomic composition, and stereochemistry calculations.
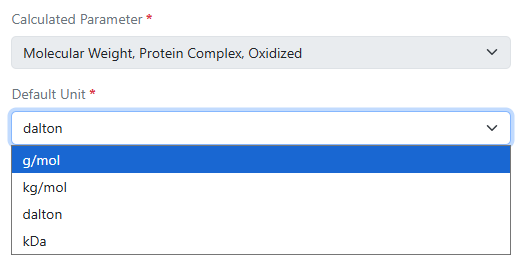
Administrators now have the ability to choose different display units for calculated molecular weight properties in DNA, RNA, Protein, and Custom material types and the preconfigured Antibodies, Plasmids, and Primers libraries. Options include g/mol, kg/mol, dalton, and kDa.
Search
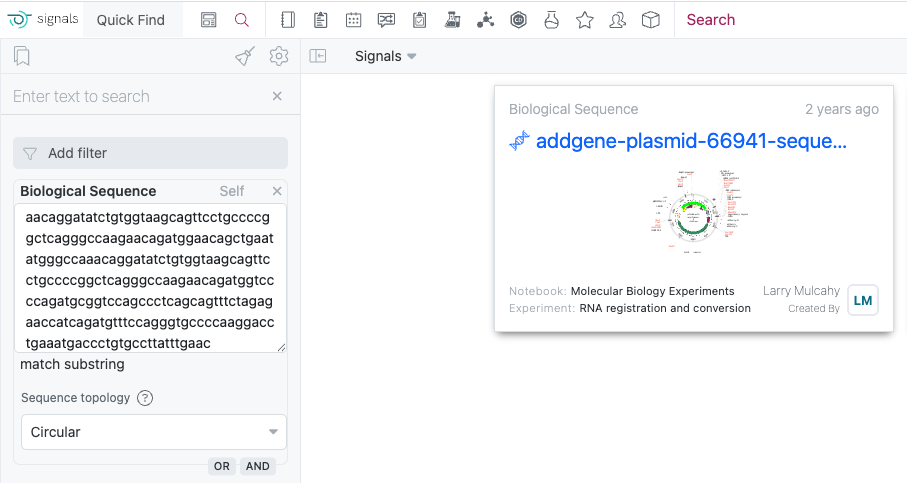
The Biological Sequence search now includes enhanced topology options to optimize your search results. The Linear option is for standard DNA and protein sequences, while the Circular option is ideal for plasmid searches where your target sequence may span across the junction between the end and start positions of the circular DNA molecule.
Administration
Signals One Private Cloud environments with Spotfire for Signals Custom Apps enabled now allow the export of Spotfire for Signals Online Debug Logs.
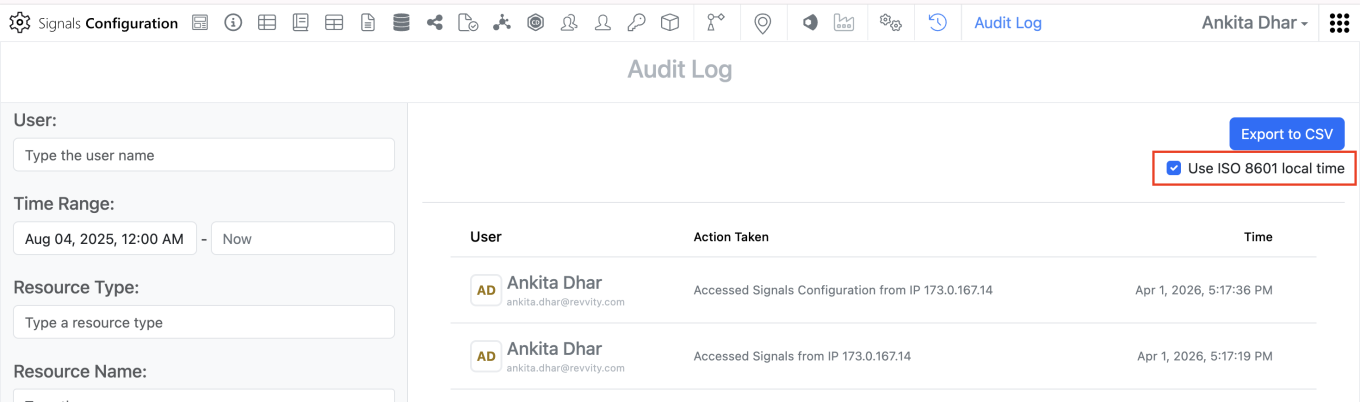
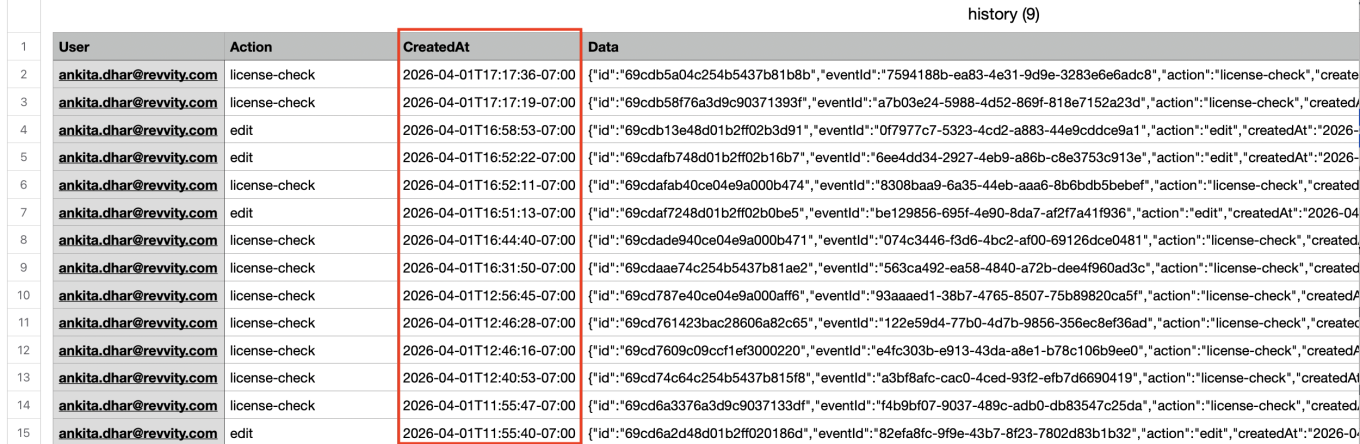
The ‘Createdat’ and ‘modifiedat’ fields in the system audit trail CSV export can be converted to local time in ISO 8601 format. When admins export the system audit trail to CSV, they can adjust the format of timestamps by selecting the “Use ISO 8601 local time” option. This setting will convert the ‘createdAt’ and ‘modifiedat’ timestamps within the CSV to local military time with a UTC offset.
Integrations & APIs
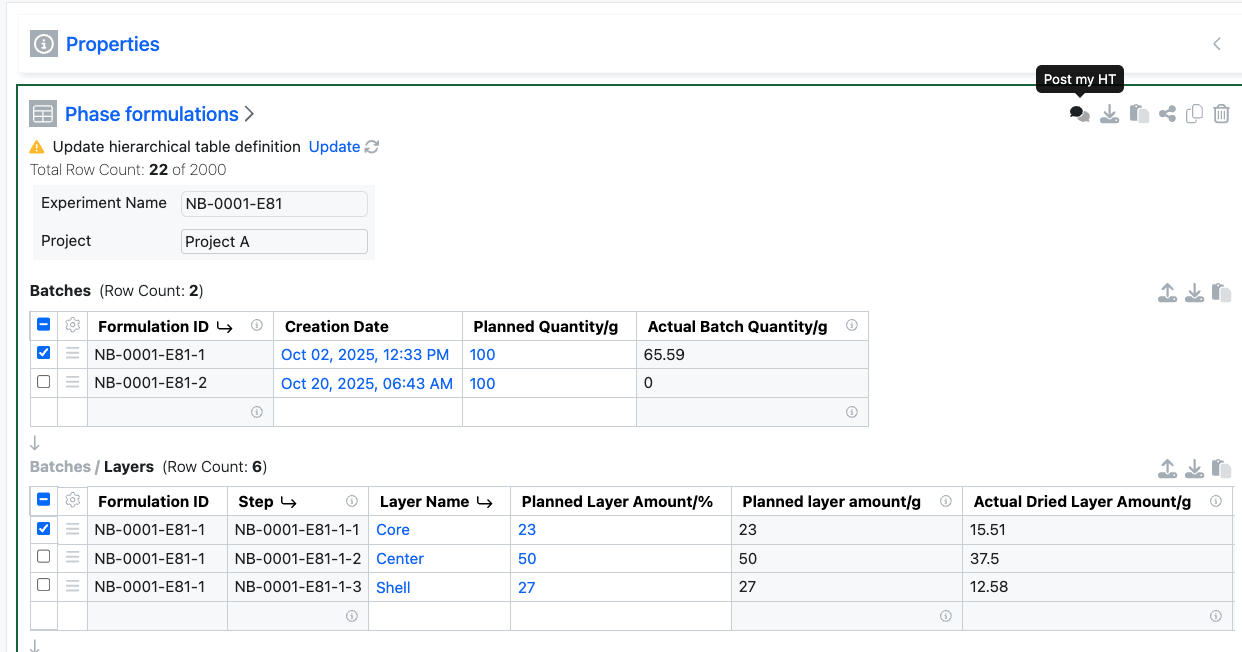
External Actions configured with a POST action on Hierarchical Tables will now include details of the selected rows and table layers within the body of the response.
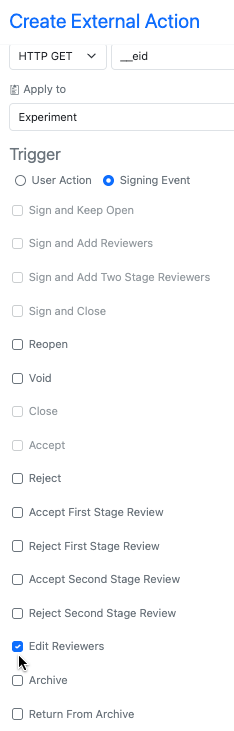
We have added a new signing event external action trigger for “Edit Reviewers” this allows for maintaining any external actions used for initially adding reviewers to also be used if those reviewers are edited.
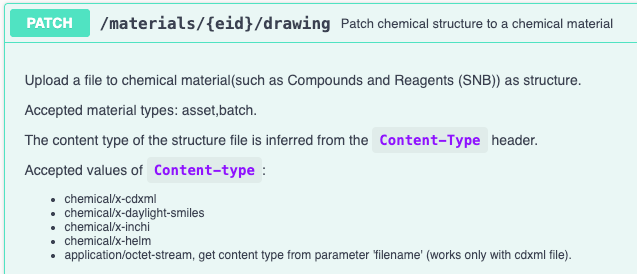

We have enhanced the API for patching material chemical drawings to accept additional chemical structure formats. The API now supports SMILES, InChI, and HELM chemical structures. You must specify the which in the request “Content-type” header. For the new content types the “filename” parameter will be ignored.

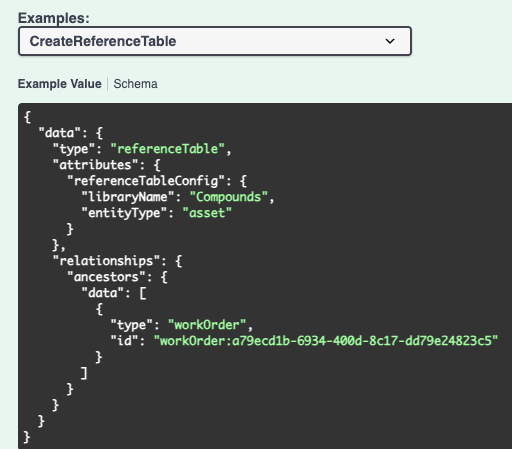
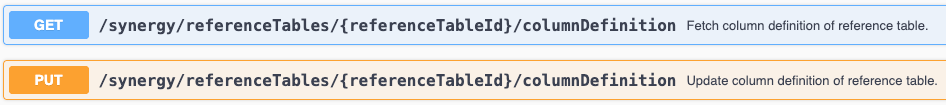
We have enhanced existing APIs to allow for creating Reference Tables in Synergy Work Orders. The library and entity type must be included in the request body. This will create a new reference table with no columns configured. Existing APIs for editing the displayed columns should be utilized to show columns.

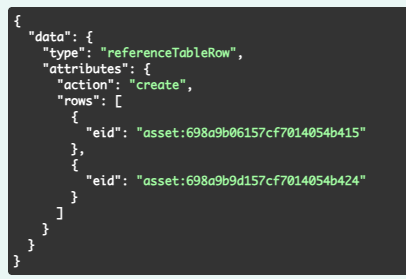
We have added new APIs to add or delete reference(s) for existing Reference Tables. You must specify the action (create or delete) and the specific entities for the rows being added or deleted.
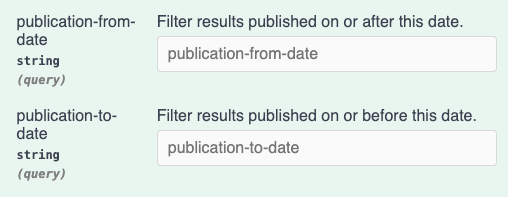
We have enhanced the search APIs for Signals Data Factory. You may now specify publication-from-date and publication-to-date as URL parameters in ISO 8601 format. This will filter results to the specified ranges.

We have added support for Proof Key for Code Exchange (PKCE) to our Bearer Token authorization flows. This is now an option for all current and future clients. This does not replace or break the Implicit Grant flow, it only adds the additional option. Additional details are available in the admin System Configuration Guide.
Beta Features
The following capabilities are in beta and are available for users, administrators and developers on the Signals platform upon request. Please contact your account representative or our support team if you would like access to the following features.
Data Workflows & Analytics
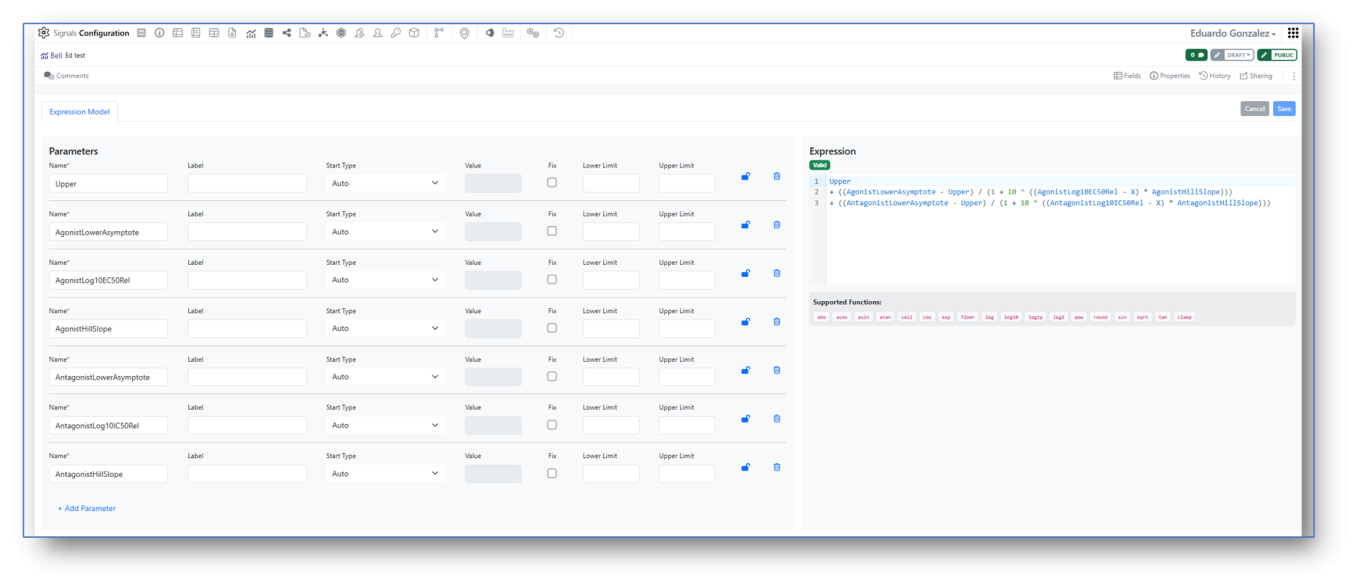
Signals Configuration allows creation of custom Analytics definitions for Curve Fitting. A mathematical Expression Editor is now available in conjunction with the settings associated to the expression parameters, which can be locked to ensure consistent start values and limits.
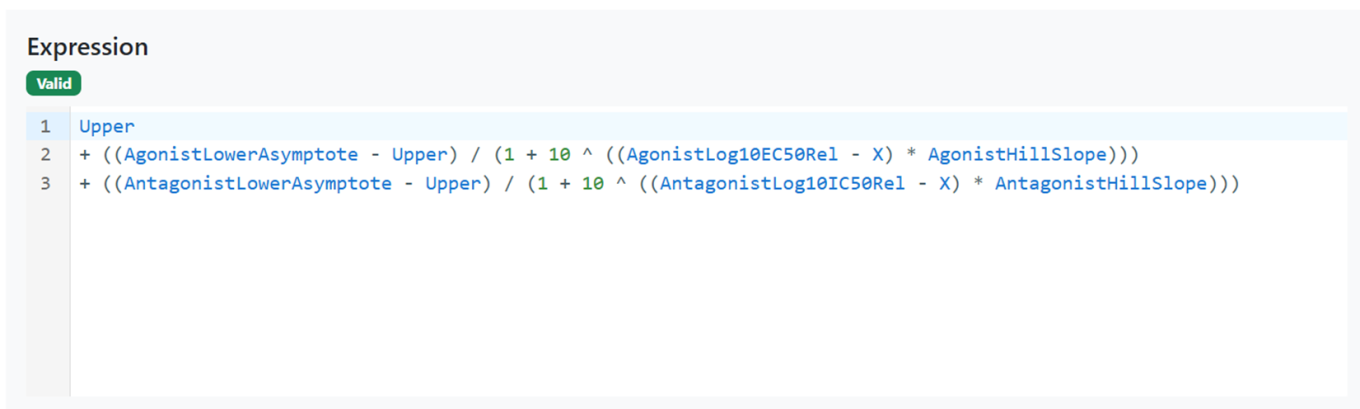
Access to the custom Analytics definitions is restricted to System administrators and Users with the “Manage Analytics Definitions” permission. The multi-line editor checks the validity of the expression that can then be saved, enabled and shared for curve fitting (here an example of a bell-shaped expression).
The custom endpoints of a curve fit can be published for subsequent SAR & Decision Making.
Tables
Configuration administrators can now give end users the ability to alter the configuration of Scatter Plots. The option to allow end users this ability is defined within the Visualization settings.
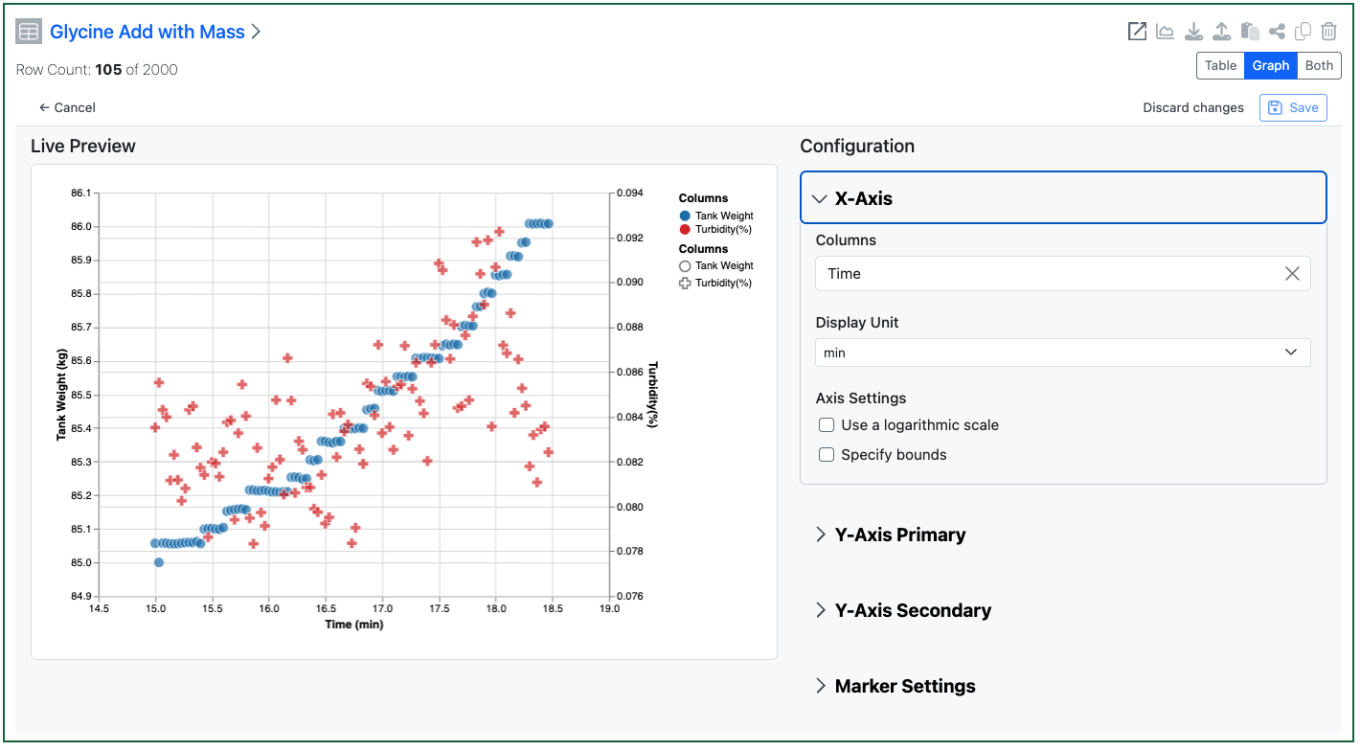
The end user can then access the setup of the chart to change the display of the axes etc. Changes are saved on demand if the user has Edit on the Experiment.
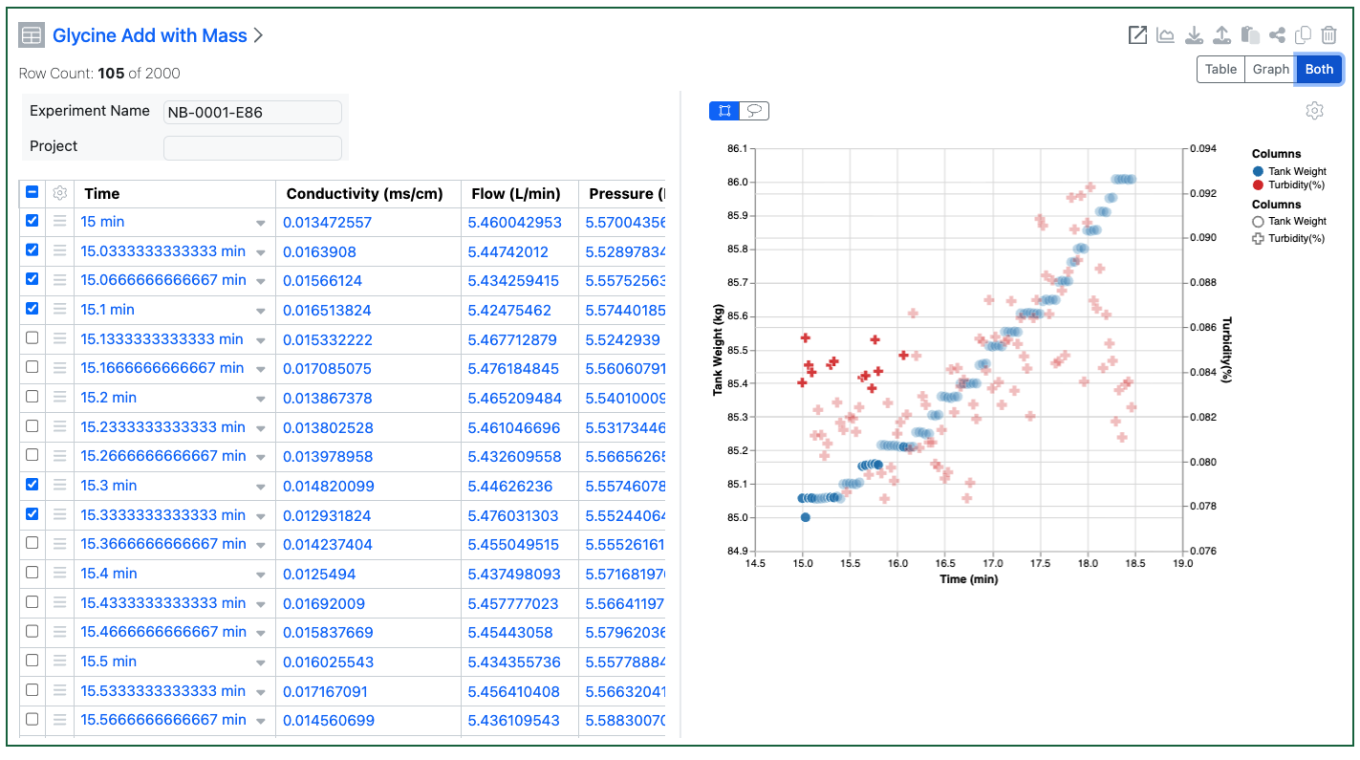
The end user can also decide to see the table and graph simultaneously. Selections in the table are highlighted in the graph view, and visa-versa. Users can select in the graph via dragging a rectangle or using a lasso to select an irregular area.
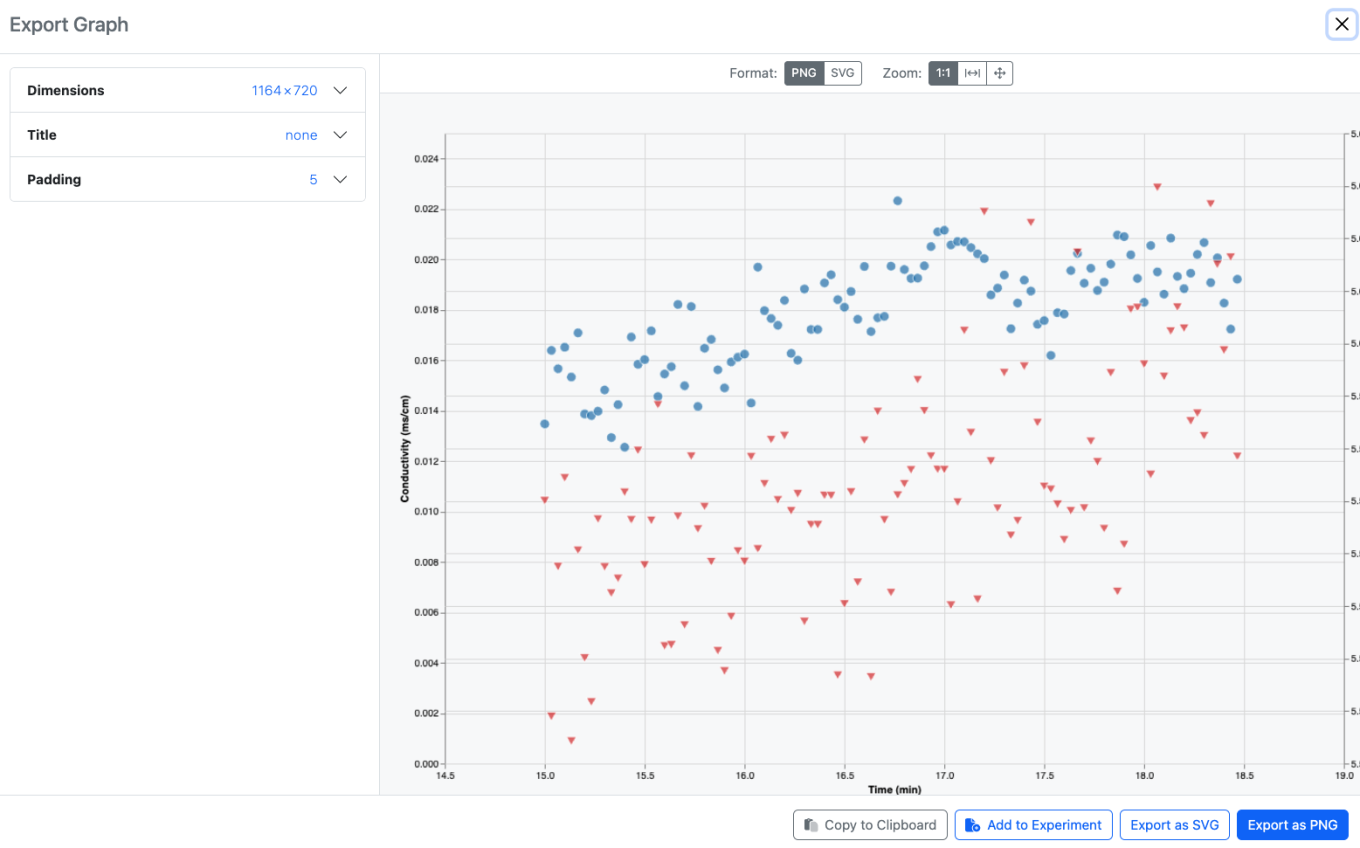
Users can also export an image of the graph, either to their clipboard, or as a downloaded file, or as an Image File directly back into the same experiment. Note that the exported image reflects the saved graph, unsaved changes by the end user will not be shown.
What's New
We are delighted to release our new version of Signals, 26.2. This release brings enhancements in Synergy, Chemistry, Materials, Inventor, SAR and Data Workflows amongst others. We have extended the grid view to the Chemical Reactions smartfolder, we have also provided a warning if users are using an outdated experiment template. We have new beta capabilities around Native Analytics and providing graphs in tables. The beta capabilities around Hierarchical tables, the Configuration Report, the updated admin experience for tables, and the View Published Data in Material libraries are now available to all systems. Finally, we have also fixed a number of small bugs.
The following improvements are available for users, administrators and developers on the Signals platform. Certain features may only be available with appropriate licensing and/or with enablement by an administrator.
- Fundamentals
– Grid View in Chemical Reactions Smartfolder
– Ontology integration now supported in Synergy, including Collaboration, Idea, Work Order, and Synergy Experiments Templates, as well as Designs and Reference Table fields (beta) - Data Workflows & Analytics*
– Grid view for normalization and curve fitting results
– Data Parser robust data type selection
– Administrative access to Signals Native Analytics (beta)
– Curve fitting extended to Logistic regression 3-4-5 Parameters models (beta) - SAR & Decisioning*
– View Published Data from Material Library is now available to all systems
– Overwrite Publications via the Publish Results to Data Factory app
– HELM Matched Monomer Pairs ChemCharts data function
– Extract GenBank Feature ChemCharts data function - Synergy†
– Advanced Search for CRO users in the CRO application
– Flexible selection of Synergy Experiment Templates by CRO users in a Work Order
– Synergy Experiment properties can inherit from Work Order properties
– Access to HELM Monomer Libraries can now be granted to CRO Departments
– Append Template to Samples for CRO users - Chemistry
– Predicted pKa calculation added to the ChemDraw Analysis panel
– Hairpins can now be generated from oligonucleotides with pendant chemical and peptide attachments
– Auto-pair tool preserves user defined hydrogen bonds when pairing complementary strands and hairpins
– Auto-pair tool now supports pairing of selected oligonucleotide fragments - Tables
– Hierarchical Tables available to all systems *
– Total row limit for Hierarchical Tables available to all systems *
– Header display settings for Hierarchical tables available to all systems *
– Define Internal or External Data Sources in Hierarchical Tables available to all systems *
– Add Inventory Columns to Hierarchical Tables available to all systems *
– Internal Reference column limited to Materials in a Material table in the same containing entity setting available to Hierarchical tables available to all systems *
– Search of Hierarchical Tables available to all systems *
– Hierarchical table data optionally included in exported PDF available to all systems *
– Updated admin experience for Admin Defined Tables and Variation Tables available to all systems
– Conditional data entry for Number properties in Admin Defined Tables and Variation Tables available to all systems
– Conditional data entry for Number with Unit properties in Admin Defined Tables and Variation Tables
– All data in a table can be cleared upon copy
– Define behavior of Headers in Admin Defined Tables and Variation Tables
– Data sources in in Admin Defined Tables and Variation Tables can be deleted
– Hierarchical Tables can be selected as the source for Contextual Lists *
– Definition and display of Scatter Plots on Admin Defined Tables (beta) * - Inventory
– Add Container Property Fields to Material Tables
– Add Parent Container field to Containers Search Table
– Copy path button next to location field
– Enable Supplier List mapping from Reactants to Material properties with auto‑populate - Materials
– Structure Preview for Materials Table
– Type in sequence registration in materials table and bulk import
– List and Attribute List fields in material IDs - Administration
– Configuration Report available to all systems
– Configuration Transfer of Hierarchical Table Templates available to all systems
– Hierarchical Tables included in Configuration Report available to all systems
– Contextual Lists included in Configuration Report available to all systems
– Warning message displays when users clone experiments that are based on outdated system templates - Integrations & APIs
– Selected rows in an Admin Defined Table can be identified in an External Action
– Selected rows in a Variation Table can be identified in an External Action
– Multiple Key:Value headers for Data Sources and Notifications
– API to fetch Security Policy
– API to add structures to Chemical Drawings with BILN
* Requires Signals One license
† Requires Signals Synergy license
We also fixed several small bugs in this release. Details of the enhancements are described below.
Administrators should be aware that we are planning to retire most of the In Vivo Apps from the Signals One App Store in Spotfire. Specifically the following Apps will no longer be available: Study Designer, Dose Preparation, Baseline Capture, Sequence Of Events, Sequence File Generator, Mass Spec, Dose Verification, Business Rules, and Data Model Designer. The PK Parameters App continue to be supported and will remain in the Signals App in Spotfire in Signals One. We encourage any customers who require assistance with this transition to contact our support organization and/or engage with our Professional Services team. This change is anticipated to happen in our next release, 26.3.
Administrators should be aware that we will be retiring the CRAIS Checker capability from Signals Notebook and Signals One. This capability, which was only available via request, was built to allow integration specifically to the Patcore CRAIS Checker database. Such workflows have been superseded by the Chemical "External Checking" capability which can be used against any chemical compliance database. Any customers still using the CRAIS checker are encouraged to update their workflows to use this capability instead. We encourage any customers who require assistance with this transition to contact our support organization and/or engage with our Professional Services team. The CRAIS Checker capability will be retired by March 31st 2026.
Data Factory administrators should note that the Publication Unique ID and Parent Publication Unique ID generation in the Publish Results to Data Factory app has been updated for robustness and now includes DateTime and the originating Experiment Name.
Data Factory administrators should be aware that the behavior of System Generated IDs has been updated. File-based publications no longer support System Generated IDs.
Data Factory Administrators should note that legacy Data Factory projects will be fully deprecated in a future release. A straightforward migration path will be provided in an upcoming release to support this transition.
Data Factory administrators should also note that the Advanced Security policies in Inventa apply an "AND" logic across all user groups, restricting access if any groups lack permissions. This behavior differs from the standard Signals security model and will be fixed to align with the standard in a future release.
Data Factory administrators should note that POST /api/v2/projects/{projectId}/results/{resultSchemaId}/rows is deprecated. Instead, supply a Unique Result ID and use PUT /api/v2/projects/{projectId}/results/{resultSchemaId}/rows/{rowId}
Administrators are recommended to subscribe to the channels within our support news site found at https://support.revvitysignals.com/hc/en-us/categories/360004446171-Support-News which contains more information about releases and other pertinent product information.
This content is anticipated for release on our Production E3 environments, and for Private Cloud customers on our deferred release schedule, in July 2026.
Further Details
The following improvements are available for users, administrators and developers on the Signals platform. Certain features may only be available with appropriate licensing and/or with enablement by an administrator.
Fundamentals
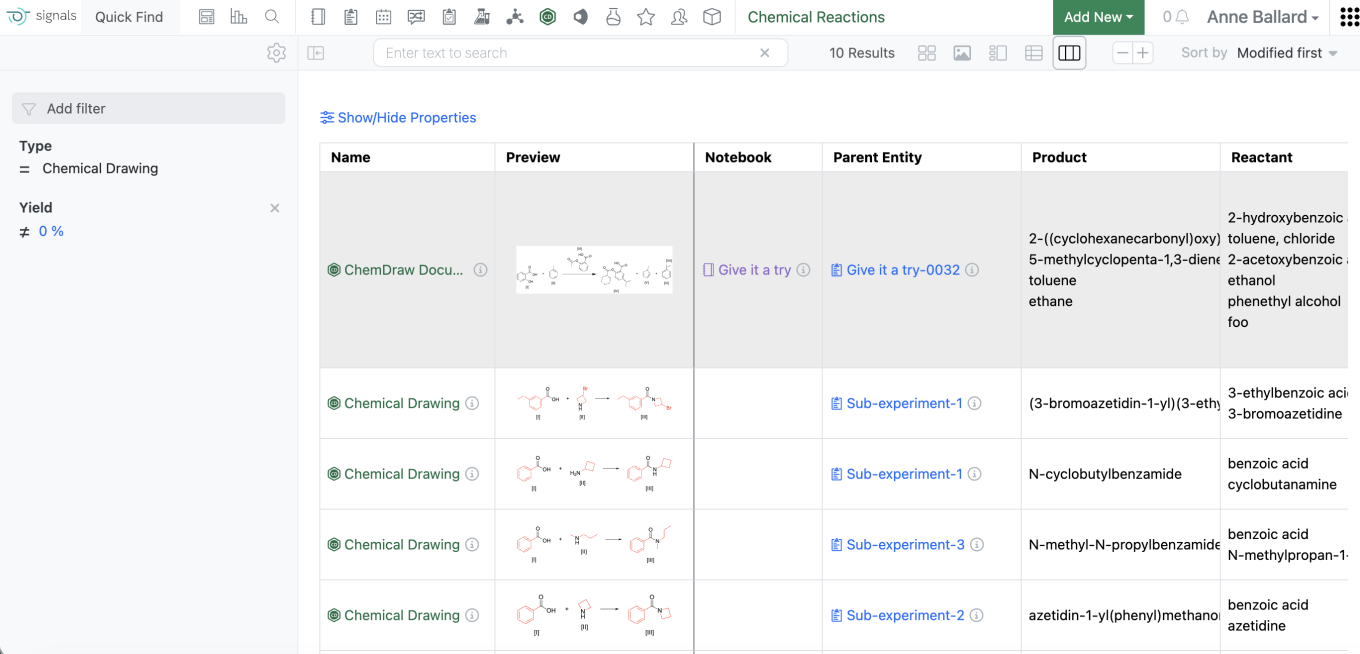
The grid view is now available in the Chemical Reactions Smartfolder. As in other grid views, customers will be able to select the properties they want displayed and can sort or filter on properties by selecting the appropriate column header. It should be noted that export is not currently available in the chemical reactions grid view at this time but will become available in a future release.
Data Workflows & Analytics
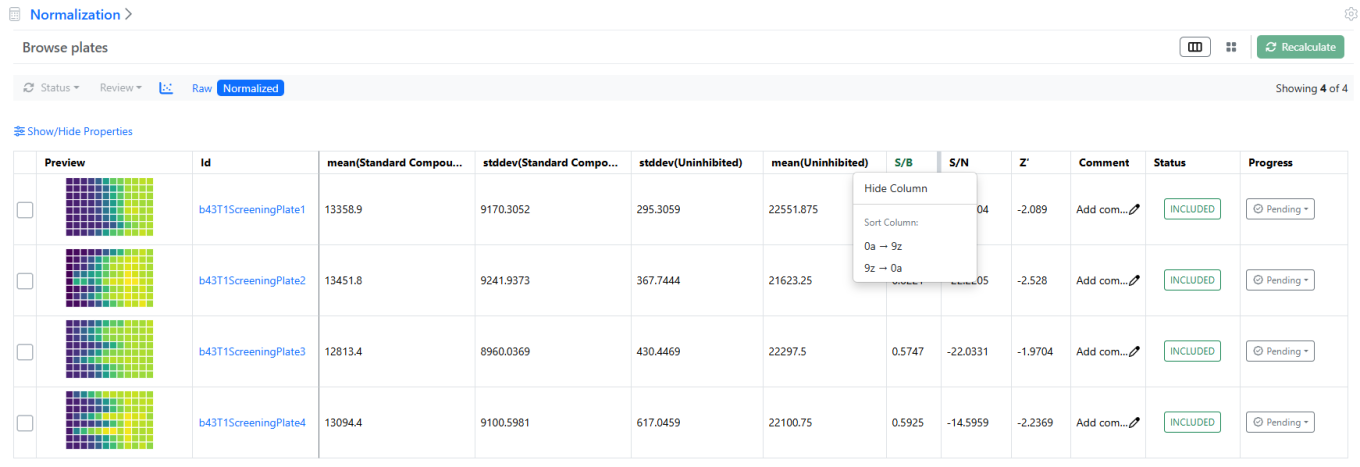
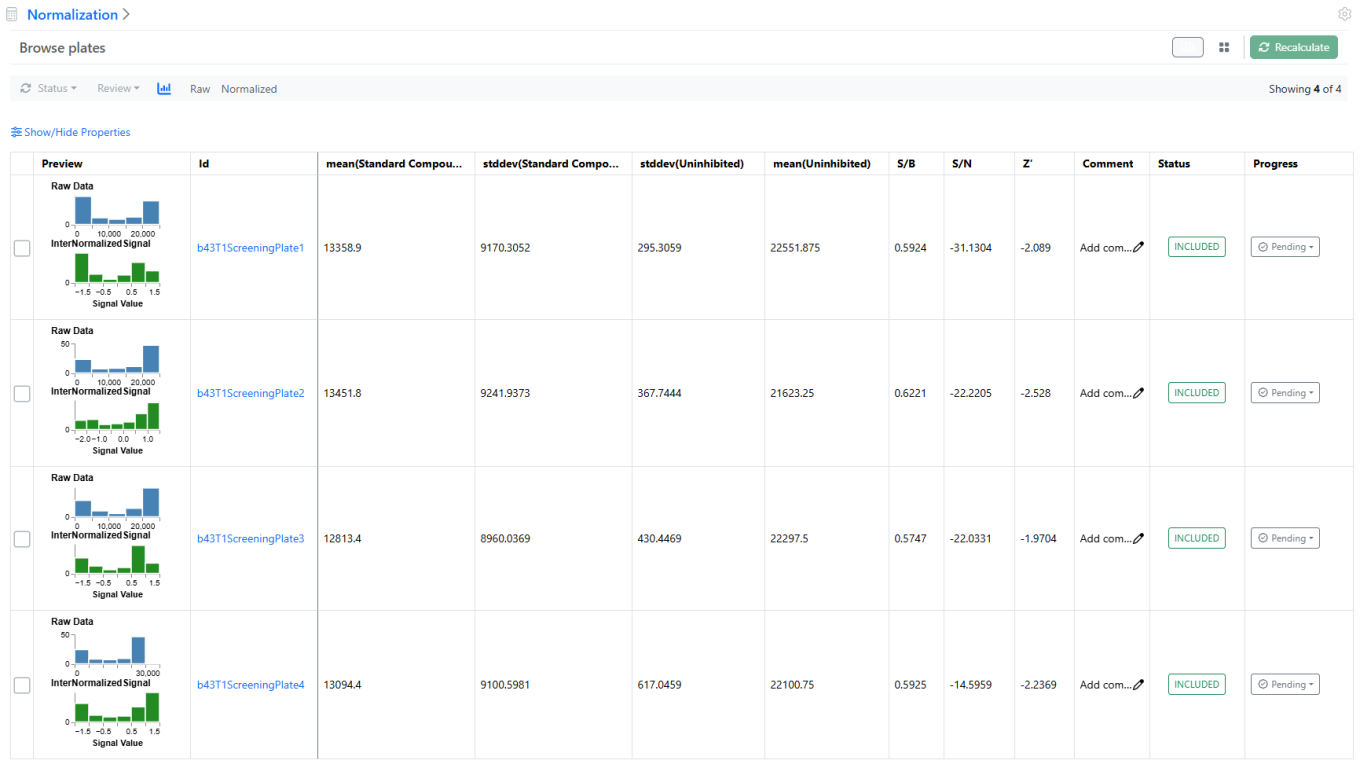
In addition to the Card view, a Grid, or Table view is now available to review both Normalization and Curve Fitting results. This tabular presentation is not only more compact but also allows hiding or reordering any column (e.g. Signals to Background ratio or IC50) to rank the results.
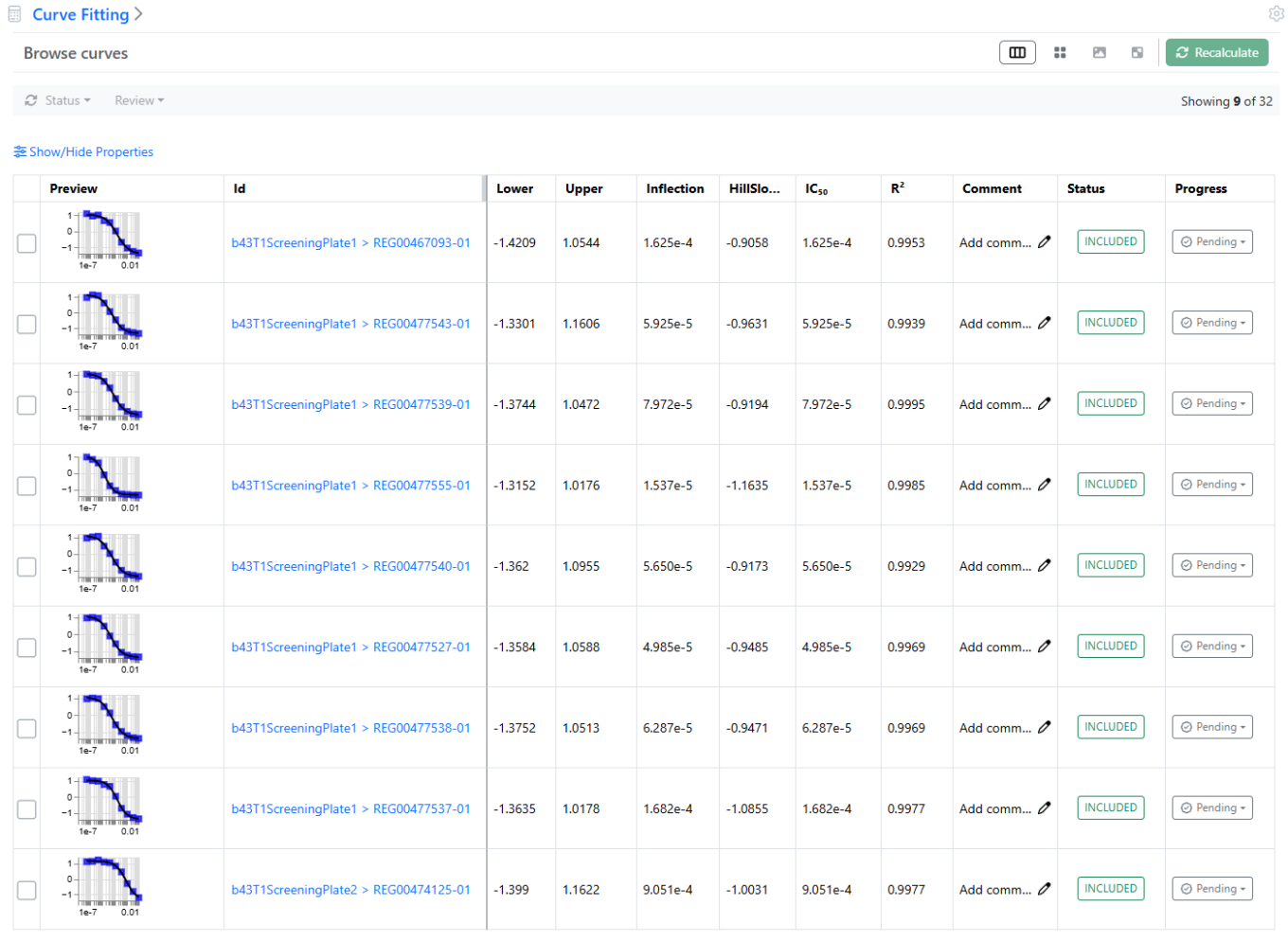
In the case of the Normalization results, the plate heatmap or the histogram views are available, as shown above.
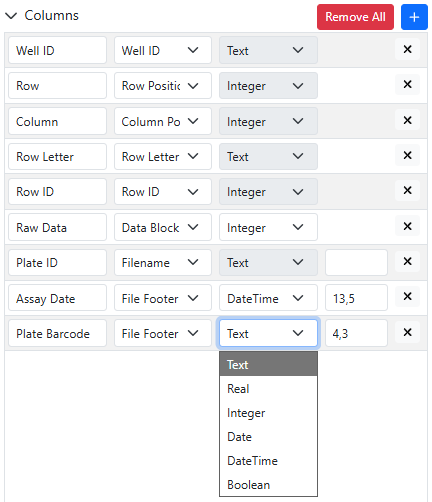
At the level of the configuration of Parser Definitions, when creating or modifying a Parser Definition, the type of data extracted from the instrument data file is either predefined (and grayed out) or can be selected from a consistent list of options.
SAR & Decisioning
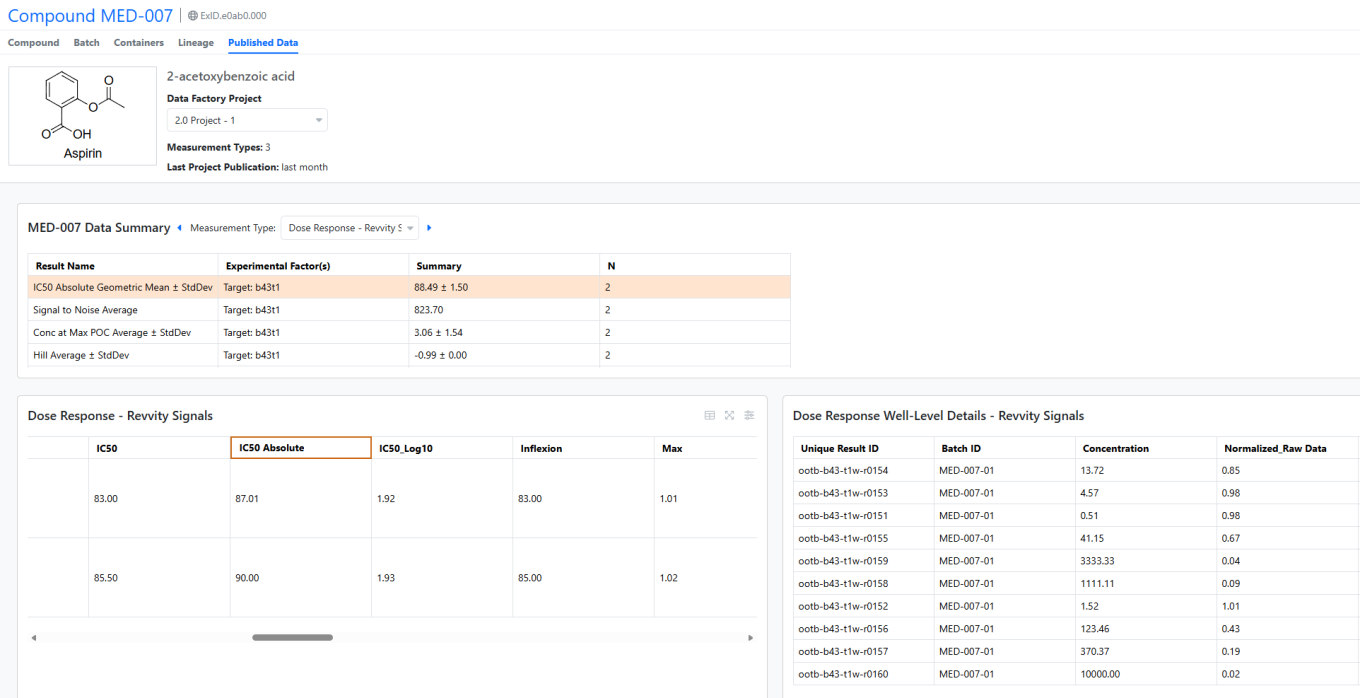
Users may now access Published Data directly from a Material Library. The Published Data tab automatically retrieves results for the selected material, displaying aggregated values by measurement type in a summary table. Interactive tables allow users to drill down to specific measurement results.
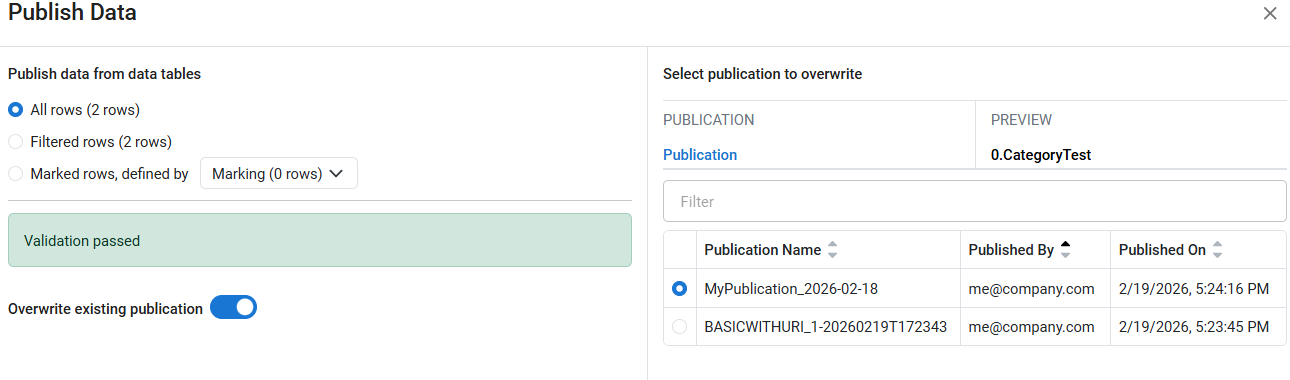
Users can now overwrite an existing publication when publishing results from a Spotfire workflow. In the Publish Results to Data Factory app, selecting Overwrite existing publication allows users to choose a prior publication, which is fully replaced with the new data. Standard users can overwrite only their own publications, while administrators can overwrite publications created by any user.
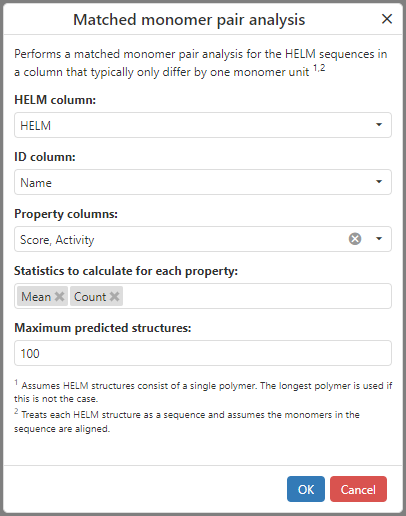
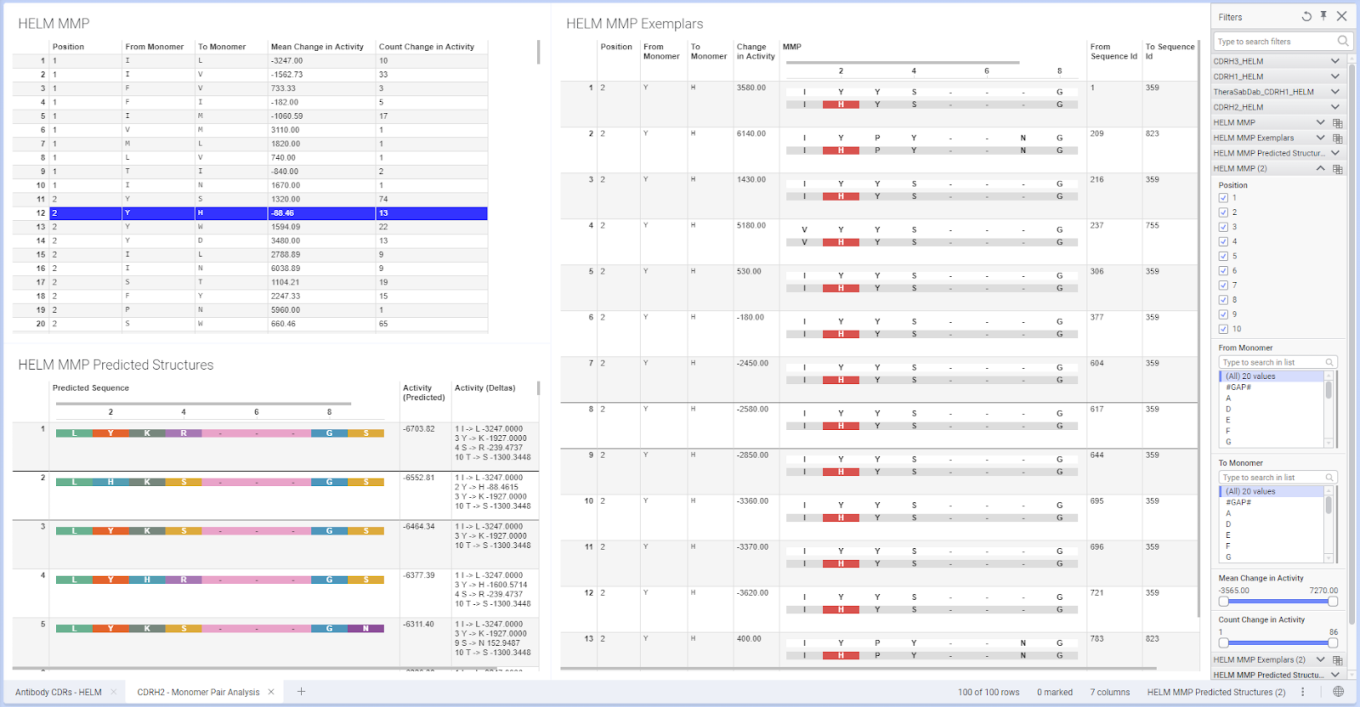
The latest ChemCharts introduces Matched Monomer Pair (MMP) Analysis for HELM sequences. HELM MMP Analysis identifies pairs of aligned HELM sequences that differ by a single monomer and quantifies how the substitution impacts numeric properties. The analysis also generates ranked predicted structures based on the learned MMP effects.
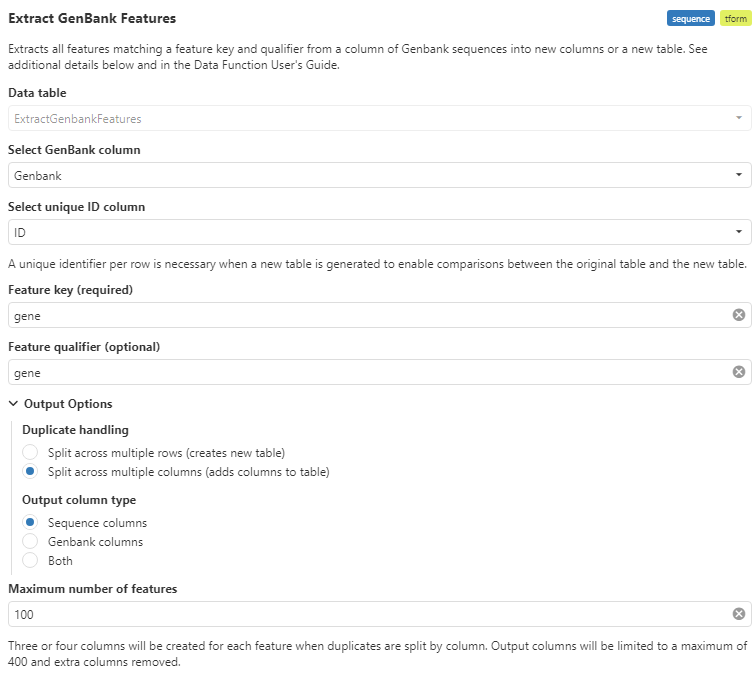
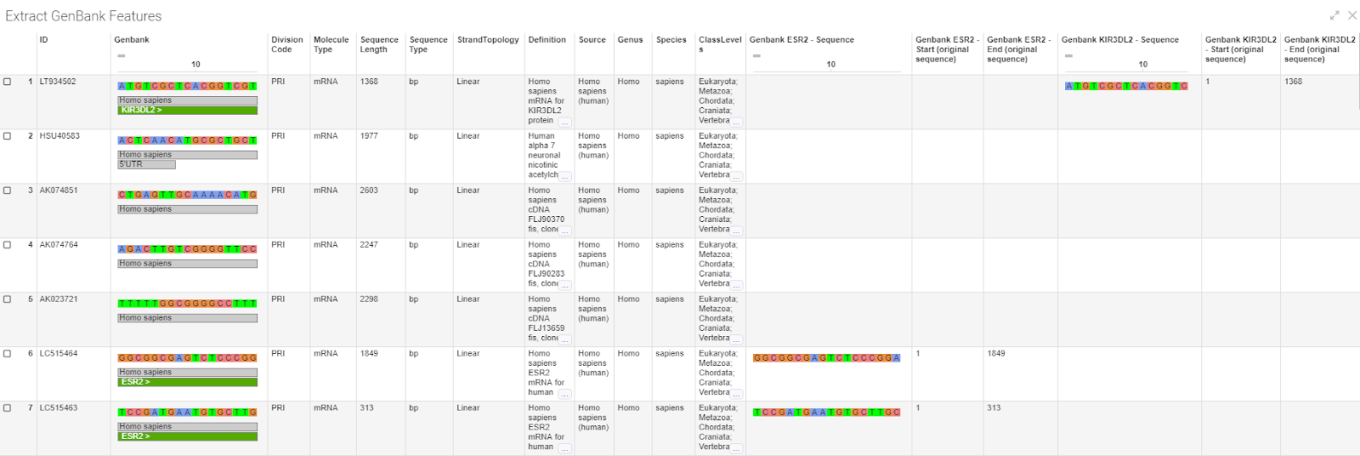
The new ChemCharts Extract GenBank Feature data function enhances and replaces the now deprecated extract GenBank Regions data function. Users can extract and split annotated regions from GenBank sequences.
Synergy
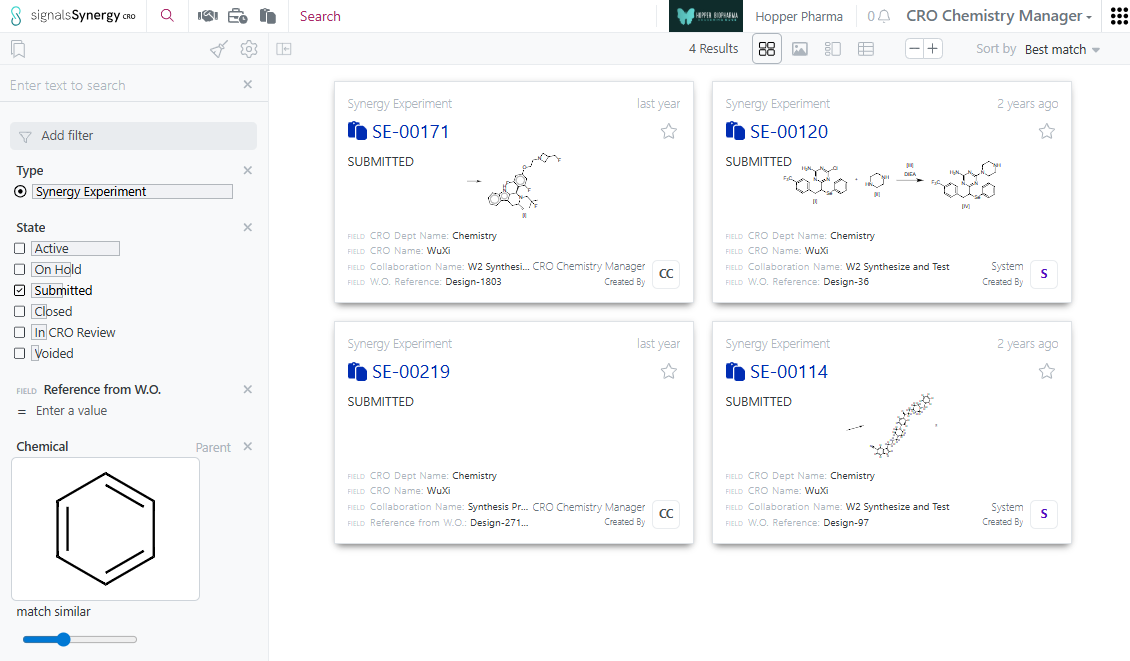
CRO users now have access to Advanced Search, including Chemical search, in the Synergy CRO application. CRO users access a dedicated search index with extra security measures, which excludes all hidden from CRO properties and ensures only entities from their CRO and Collaborations.
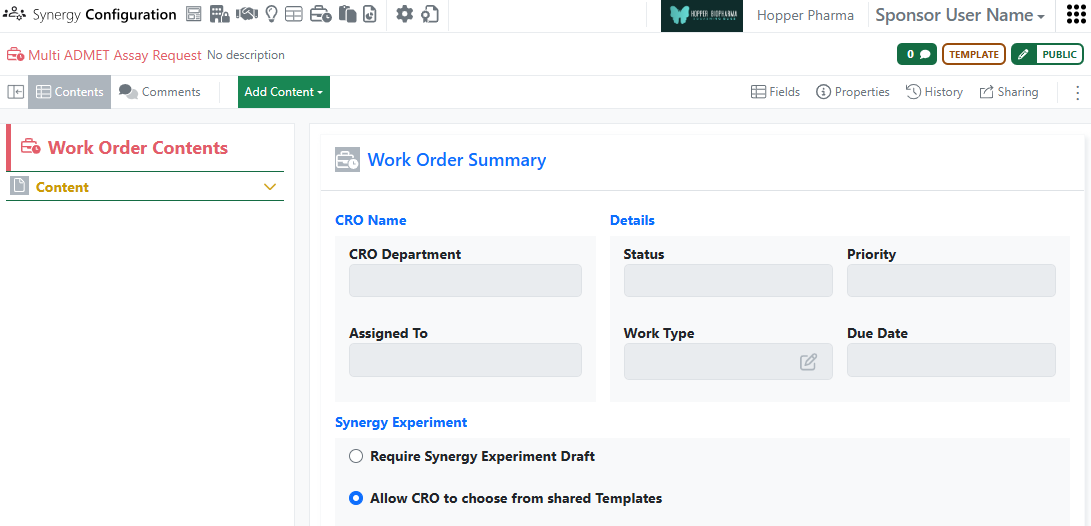
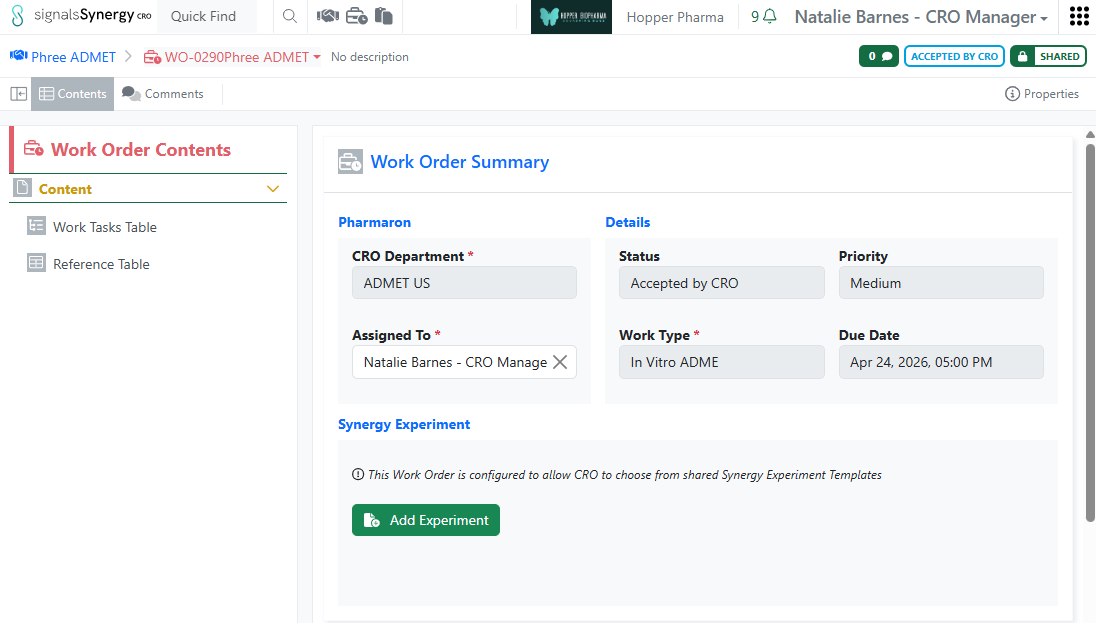
When the Sponsor wants to give a CRO flexibility in how they report their data or when the work requested requires varying experimental formats, it may be easier to allow the CRO to select from a set of shared Synergy Experiments rather than specifying a single Synergy Experiment Draft. This is especially useful when requesting multiple assays in one Work Order. On a Work Order Template, an administrator can now choose between Require Synergy Experiment Draft or Allow CRO to choose from shared Templates. When the Work Order is configured to allow the CRO to choose, no Draft Synergy Experiment is required from the Sponsor and the CRO can select from a list of shared Synergy Experiment Templates when adding a Synergy Experiment. Note that Synergy Experiment Templates are shared on a per CRO Department basis in Synergy Configuration > Synergy Experiments > Templates. Finally, when a CRO user creates a Synergy Experiment from a Template, all Hidden from CRO field values must be provided on the Synergy Experiment Template or configured to inherit from the Collaboration or Work Order.
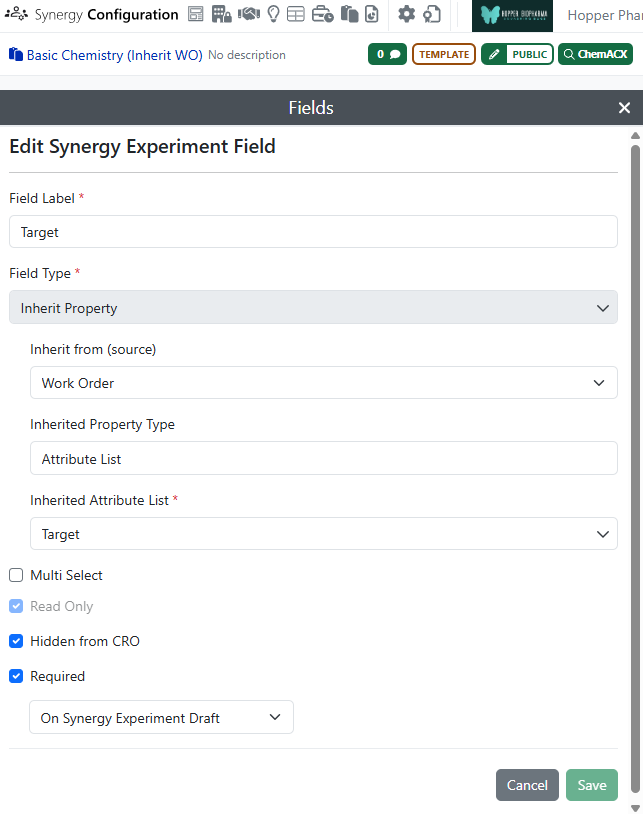
Synergy Experiment Template attribute list fields can be configured to inherit the property value from the associated Work Order. (Fields can also inherit from the parent Collaboration.) If a field is set to inherit from Work Order, the field will automatically populate with the corresponding value from the associated Work Order. This value is populated either on the Draft Synergy Experiment or when the Synergy Experiment is created from a Template. All subsequent Synergy Experiments created by CRO users will also inherit that value. As a result, the Sponsor or CRO user does not have to repeatedly populate the same property, such as Project Code or Disease Target. Further, Hidden from CRO fields can be populated when a CRO is allowed to select their own Synergy Experiment Template.
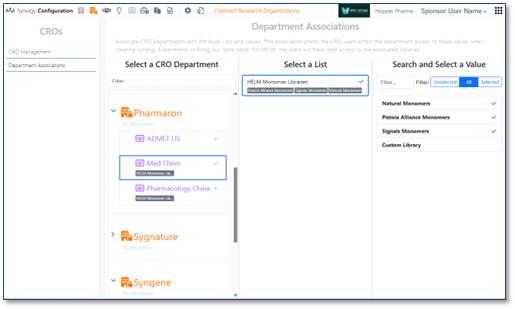
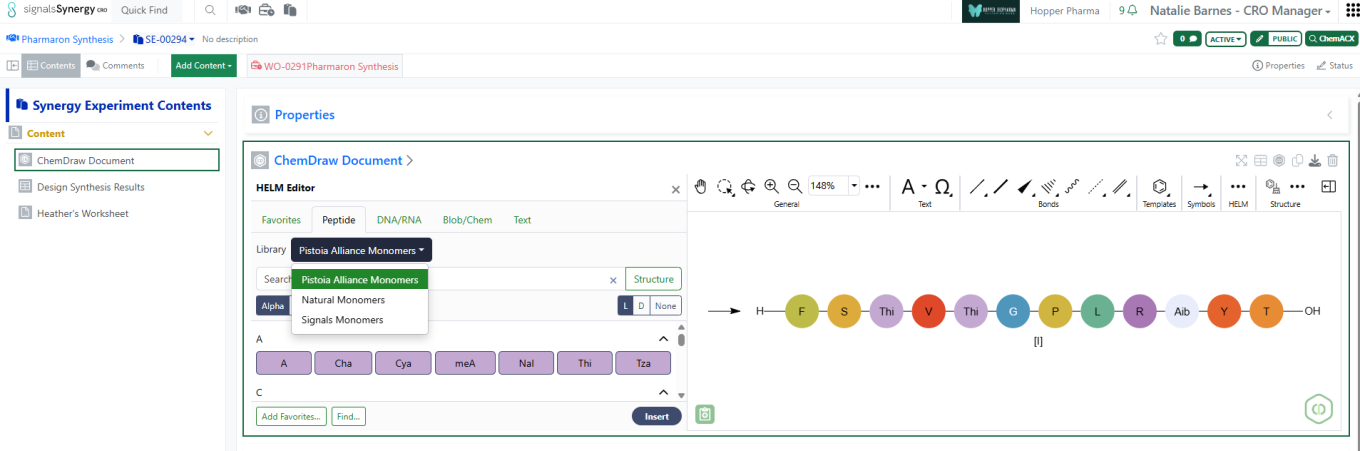
Administrators with the privileges Administer Synergy, Manage CROs, and Manage Attributes can now grant CRO Departments access to specific HELM Monomer Libraries. In Synergy Configuration > CROs > Department Associations, administrators can associate one or more HELM Monomer Libraries with a CRO Department. Users in CRO Departments have access to a HELM Monomer Library if and only if they have an association with that library. (No additional configuration is required.) With library access, a user can access to the HELM monomer editor and search the library to add monomers to their chemical drawings. (Note: As always, all users, whether internal or CRO users, can hover to see the structure represented by a monomer notation added to an Experiment, Work Order (e.g., Designs), Synergy Experiment, etc. by another user, independent of their access to a HELM monomer library.)
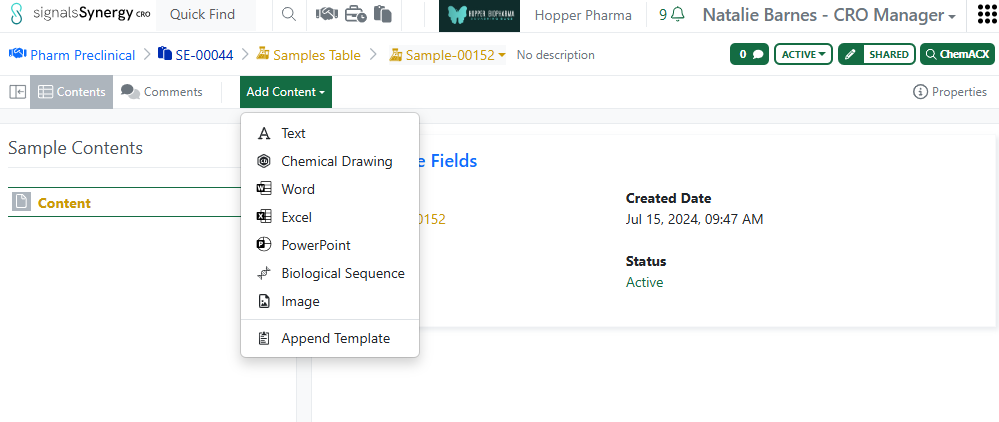
Finally, the ability to Append Templates to Samples has been extended to CRO users in Synergy Experiments.
Chemistry
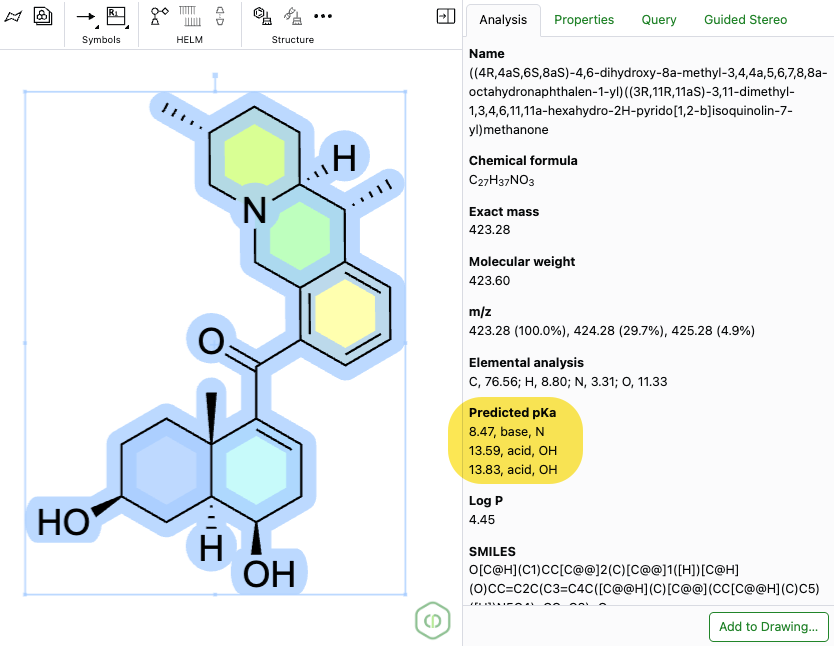
Predicted pKa is now available as a calculated property in the ChemDraw editor Analysis panel. The calculation uses the current selection, supports structures with fewer than 100 heavy atoms, and returns the predicted pKa value, type (acid or base), and atom label. When atom numbering is enabled in the drawing, the corresponding atom number is also shown in the Analysis panel.
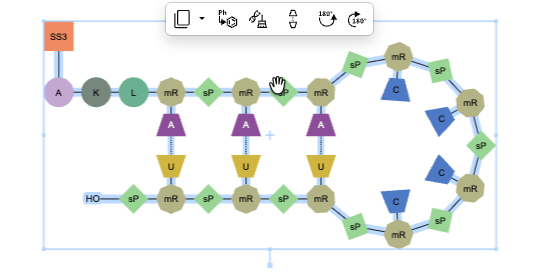
Hairpins can now be generated from oligonucleotide sequences that include pendant chemical linkers or peptides by using the auto-pair tool to automatically find base complementarity and place hydrogen bonds to form the hairpin structure.
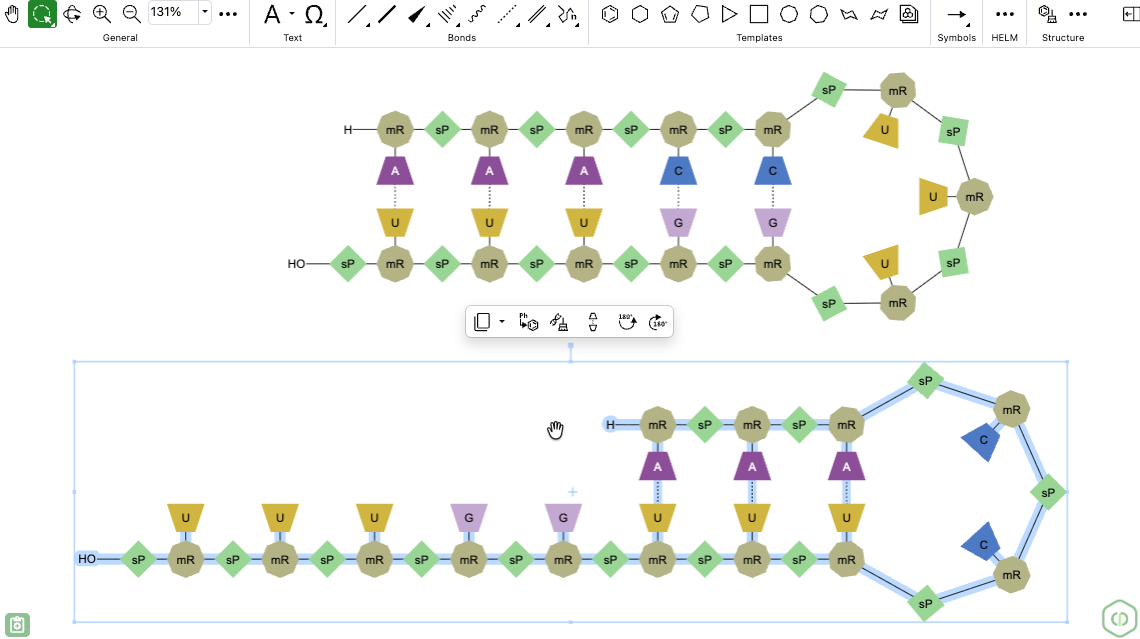
When there are multiple ways to create a complementary strand or hairpin, the auto-pair tool can now be used after placing one or more hydrogen bonds between nucleobases. ChemDraw then uses those existing hydrogen bonds as a guide, preserves them, finds the best remaining base complementarity, and adds the hydrogen bonds needed to complete the complementary strand or hairpin.
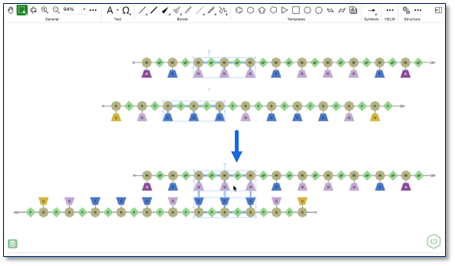
The auto-pair tool now supports partial selections of oligonucleotide strands, pairing compatible nucleobases within the selected regions and adding the corresponding hydrogen bonds between them.
Tables
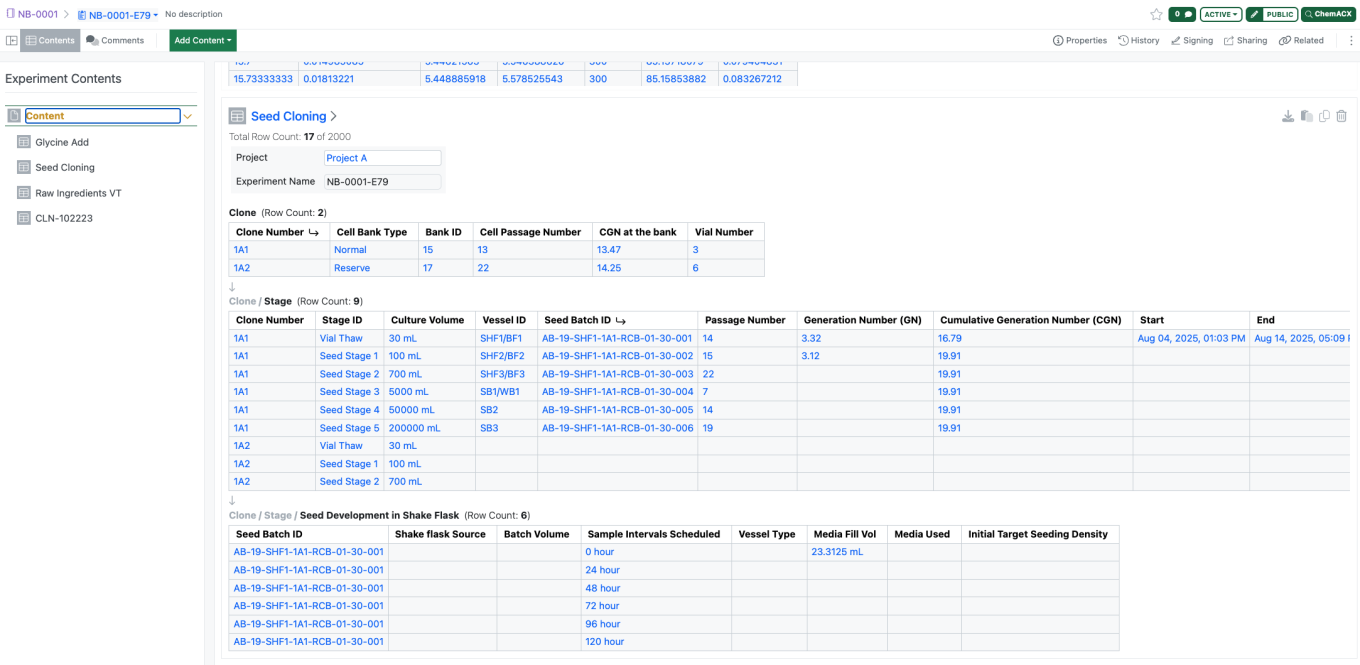
Hierarchical Tables are available to all systems.
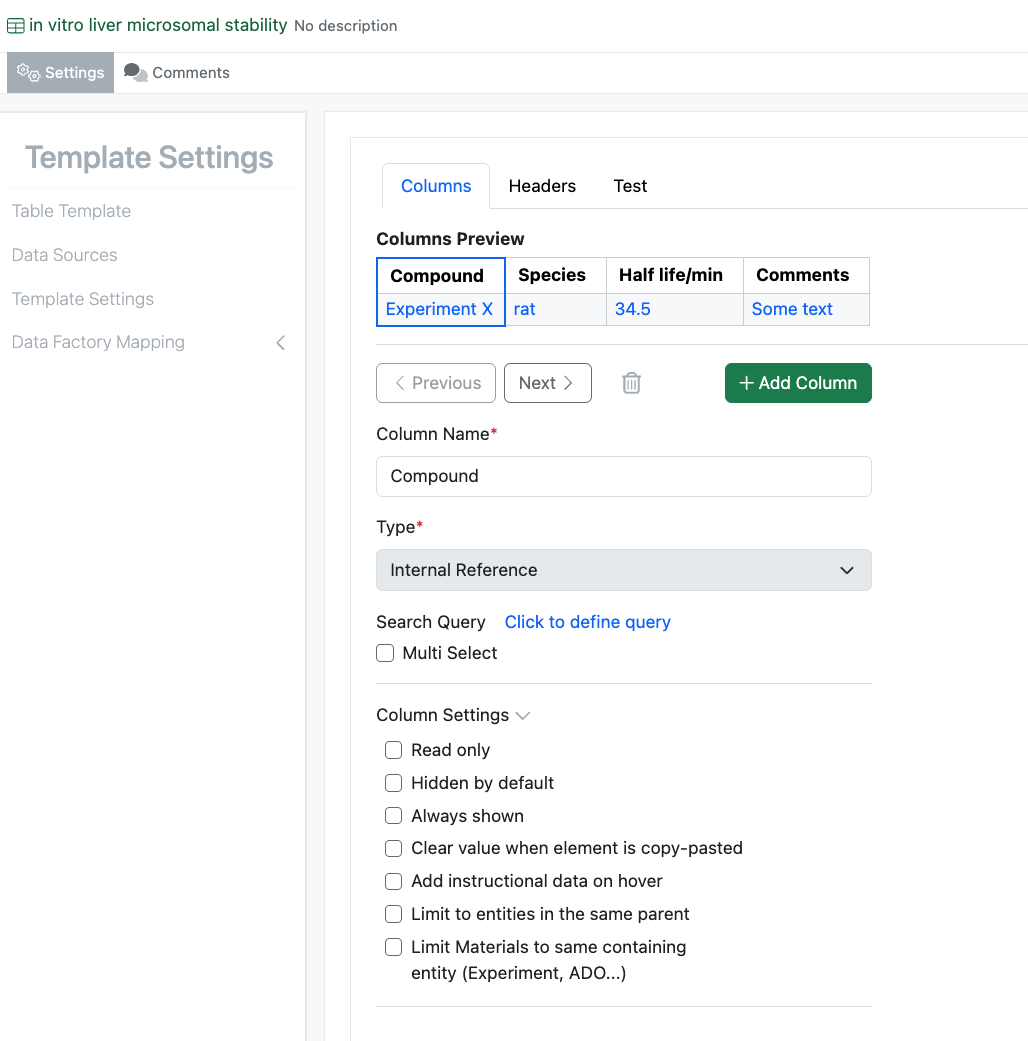
The updates to the administrative experience recently released in beta is now available for all systems.
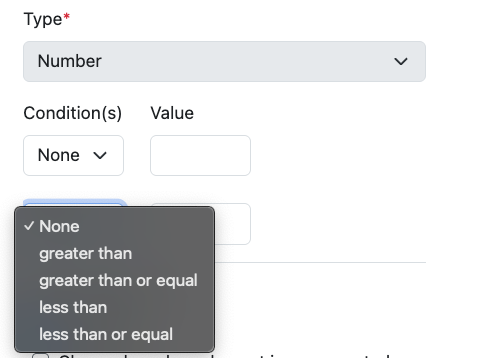
The update to allow Conditional Data Entry on properties of type Number is also available for all systems.
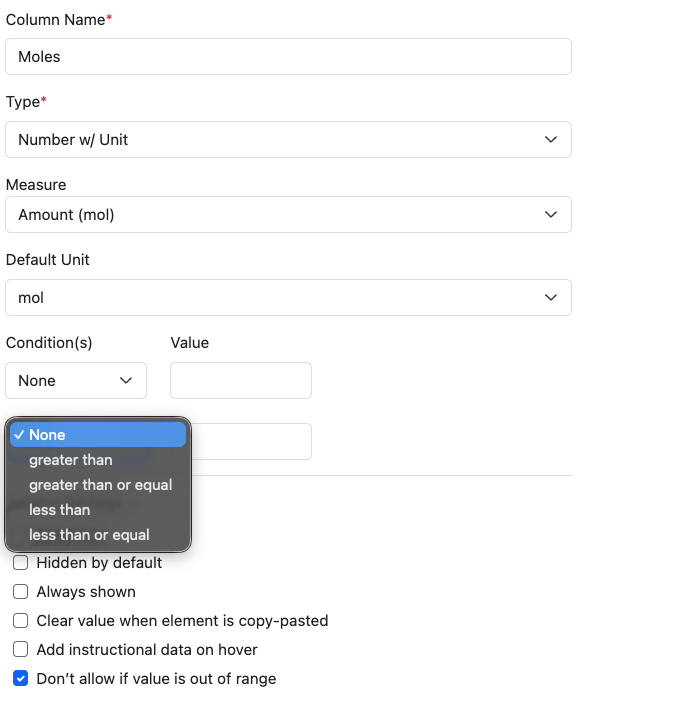
This capability has also been extended to allow Conditional Data Entry on properties of type Number with Unit in Admin Defined Tables and Variation Tables. Number with Unit properties can be defined with conditions for data entry, ranges can be set and the property set to either not allow an out of range value, or to provide a visual indication when a value does not match.
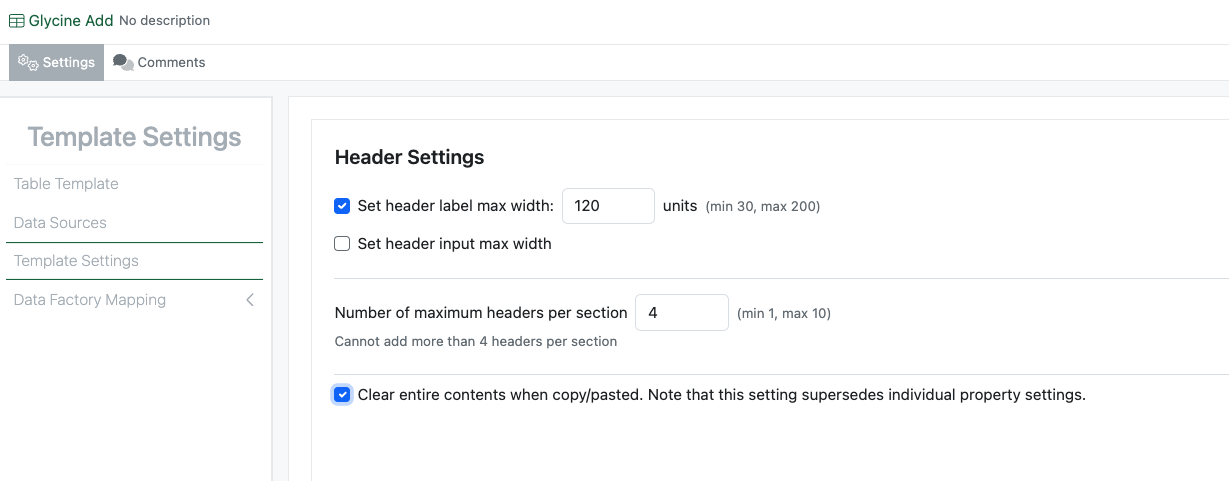
The administrator can now define that all of the data in an Admin Defined Table, Variation Table or Hierarchical Table is cleared when the table is copied. This setting supersedes any individual settings on the individual properties.
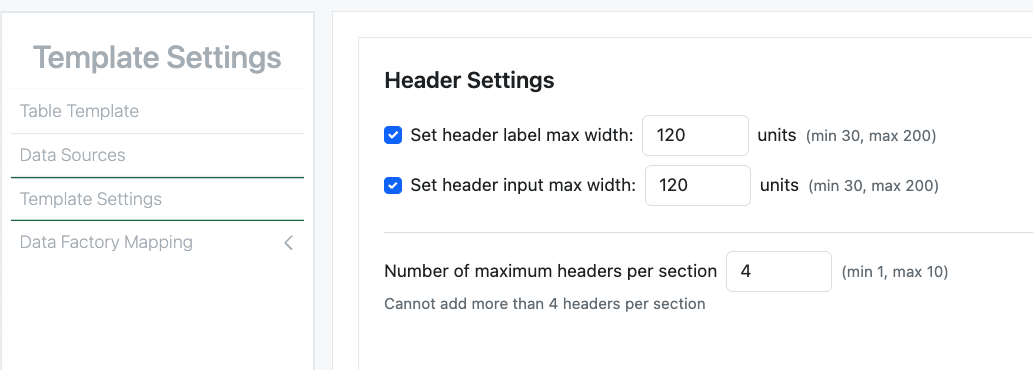
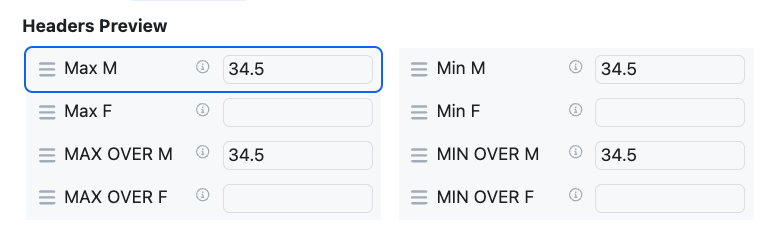
The administrative and end user experience for the definition and display of Headers in Admin Defined Tables and Variation Tables has also been updated to match Hierarchical tables. Headers are displayed in more consistent vertical array. The admin can define the maximum width of the header label, and the data entry fields, to allow optimal alignment. Admins can also define the maximum number of Headers in a vertical section, the Headers are arranged top down in a given section up to the maximum number upon which a new section is started.
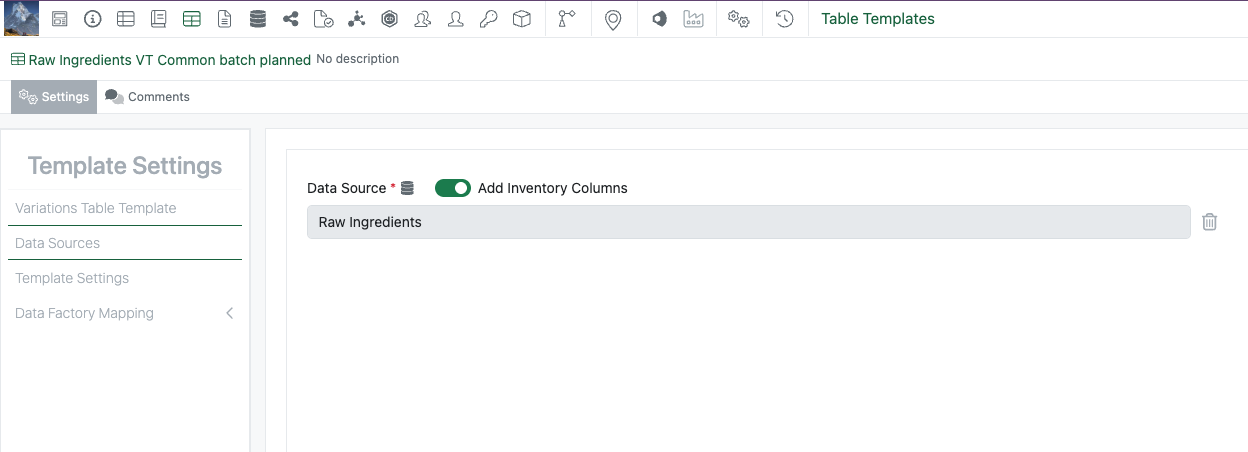
Data sources defined in Admin Defined Tables and Variation Tables can now be removed and replaced. The data source can be removed via the Data Sources panel.
Properties in Hierarchical Tables can now be used as a Source for Contextual Lists.
Inventory
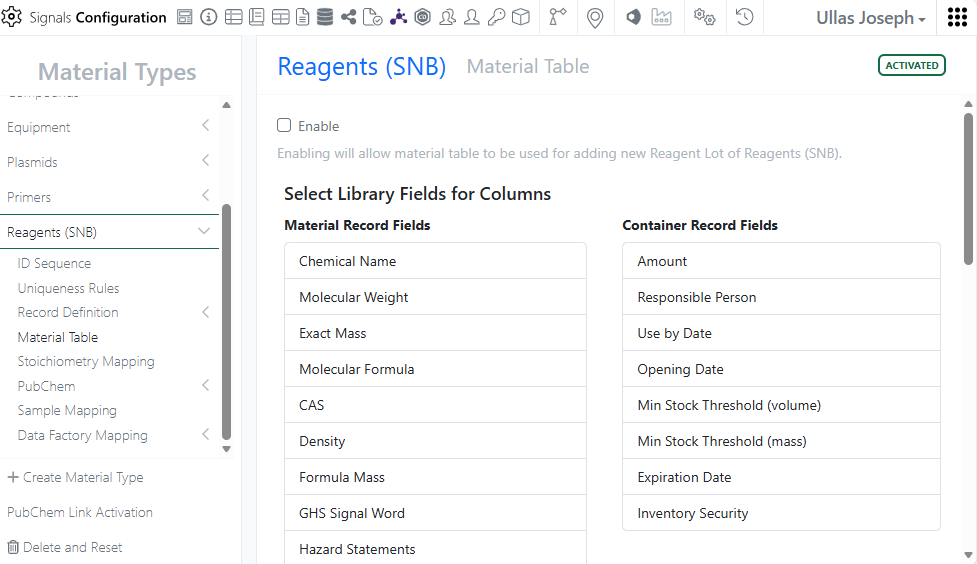
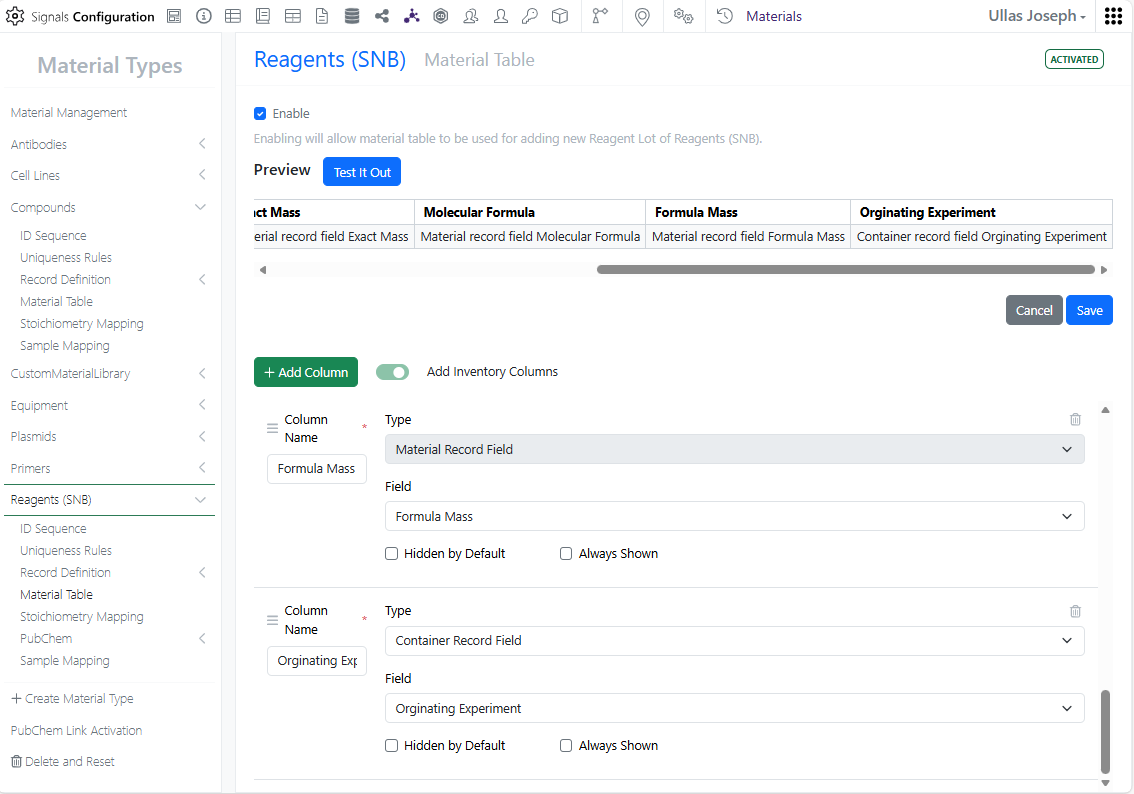
Inventory Admins can now view and add container-specific properties directly in material tables for better experiment decisions and accurate inventory tracking.
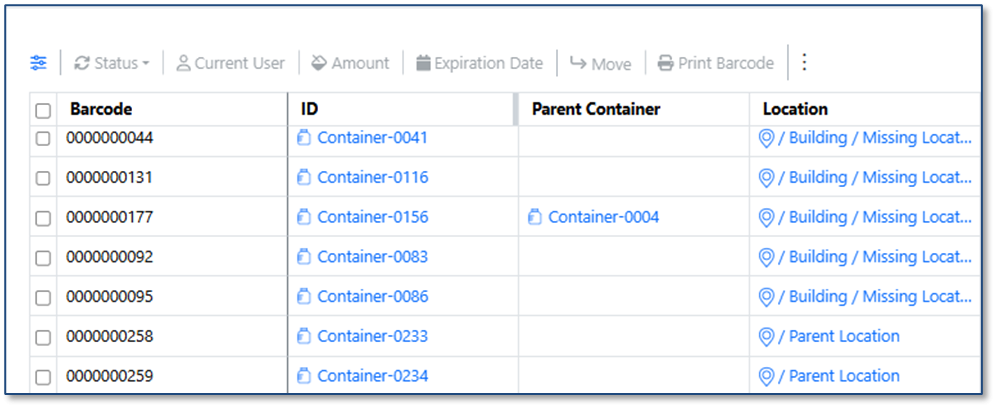
The Containers Search Table now displays the Parent Container field, making it easier to identify aliquoted container relationships.
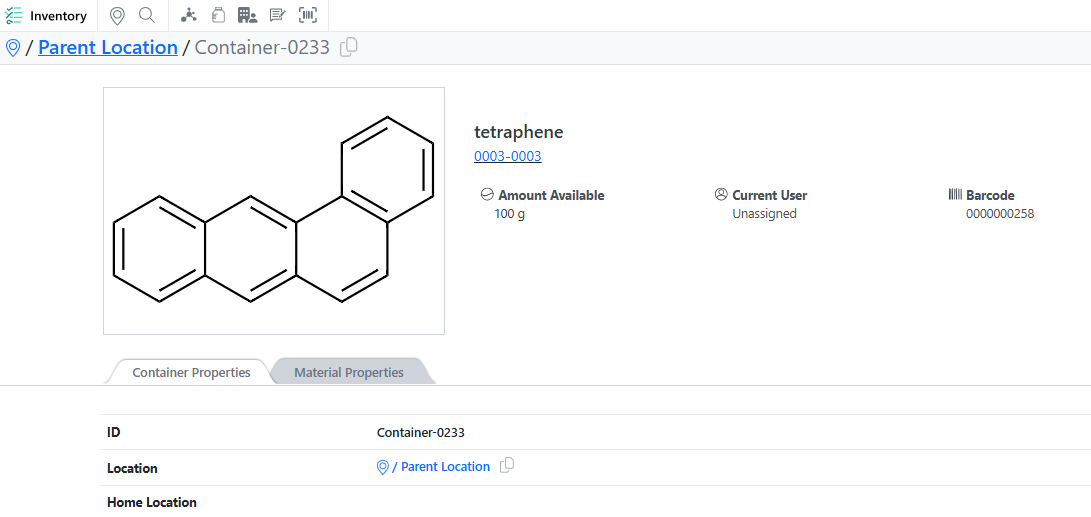
A copy button is now available next to the location path, allowing users to copy the full path as a single line.
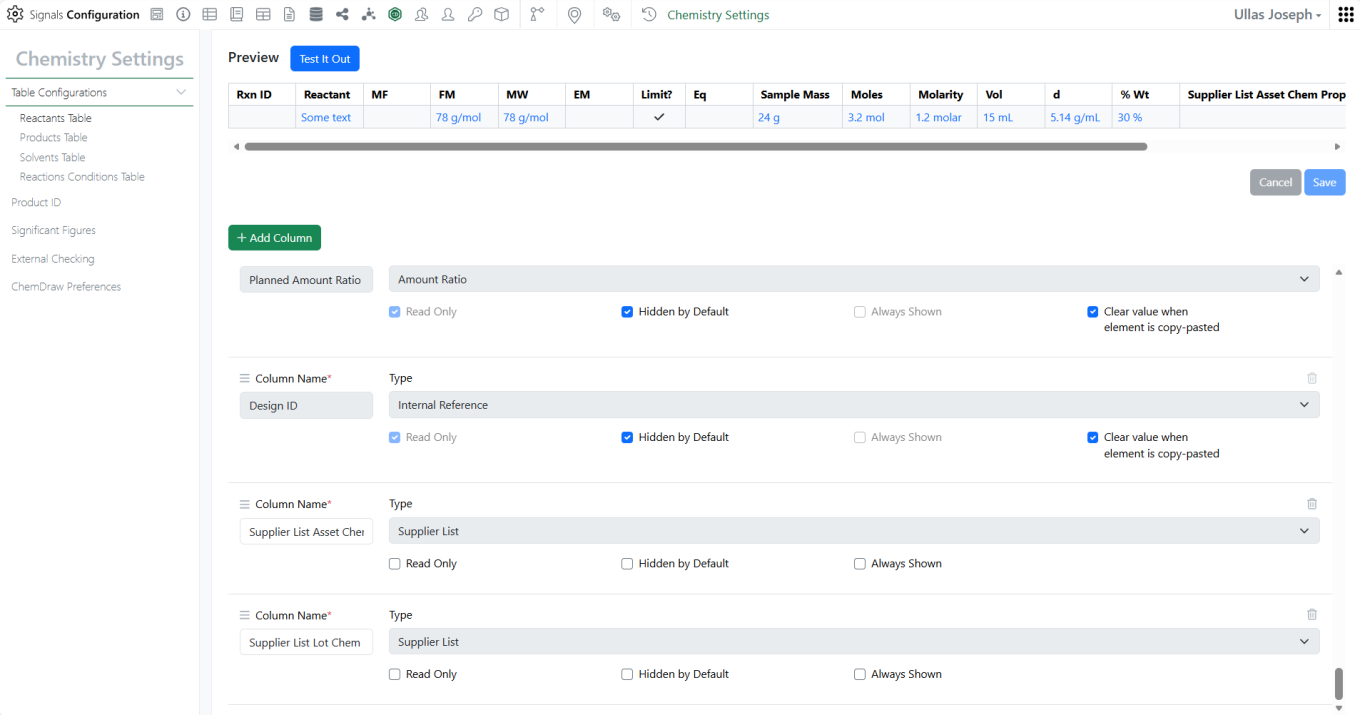
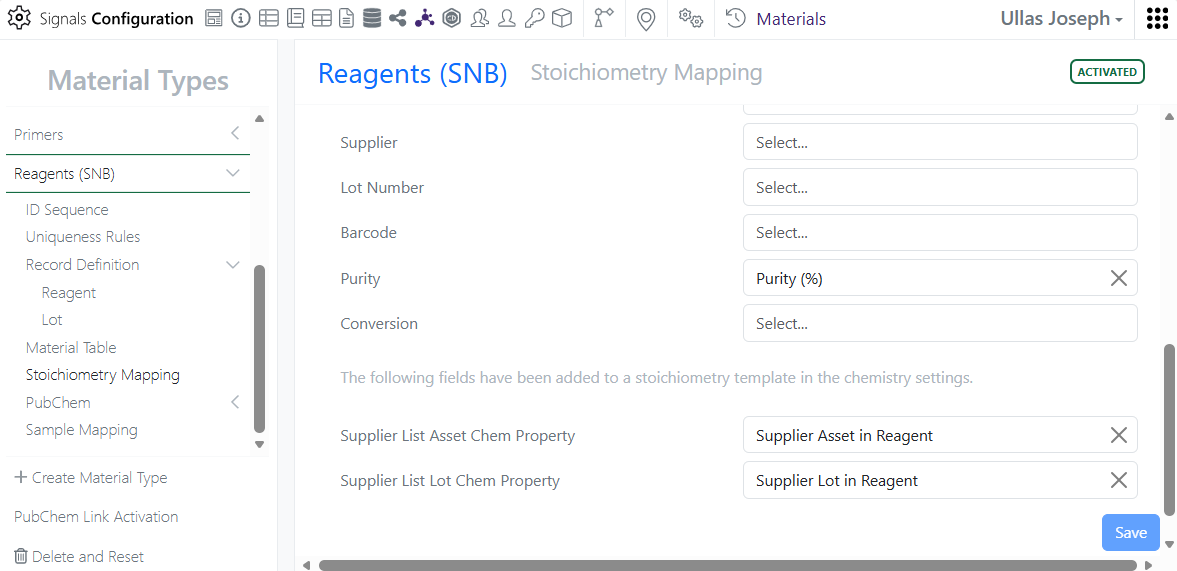
Supplier List can now be mapped from Reactants to Material Properties, auto-filling the Material field in the Stoichiometry grid.
Materials
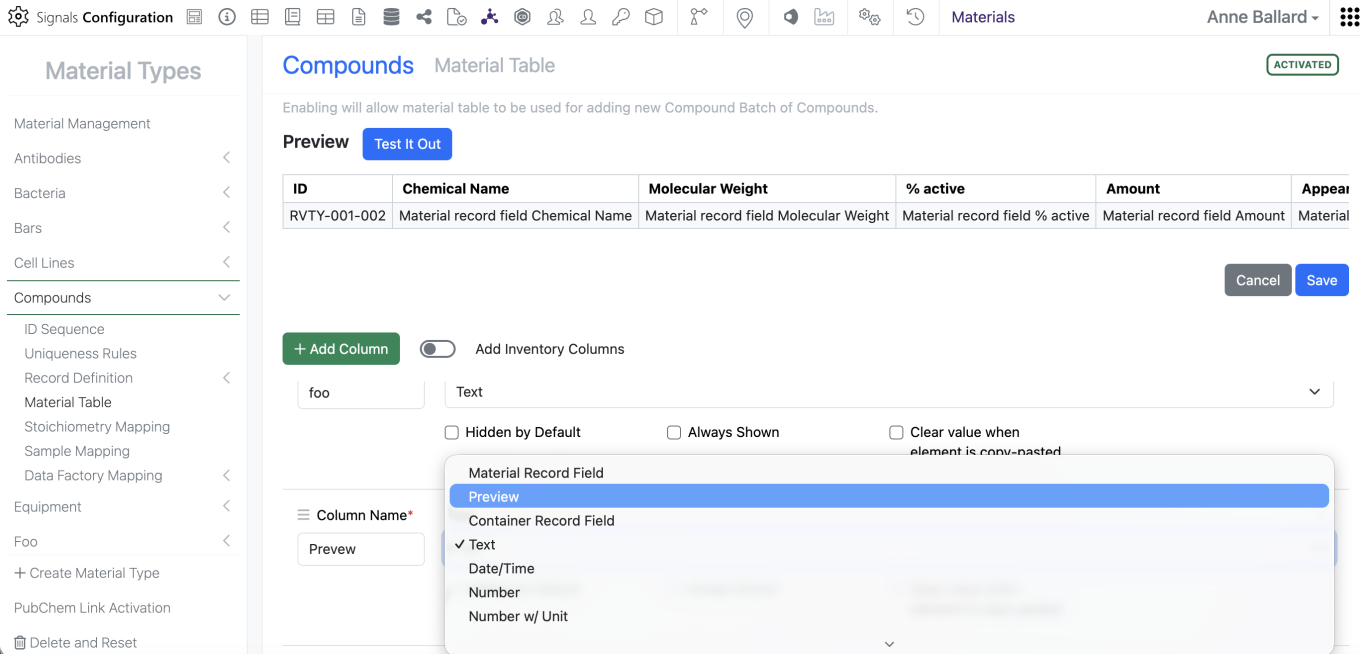
Admins are now able to add the Structure preview to the Materials Table by adding a new column of type “Preview” to the Materials table in the SN Config
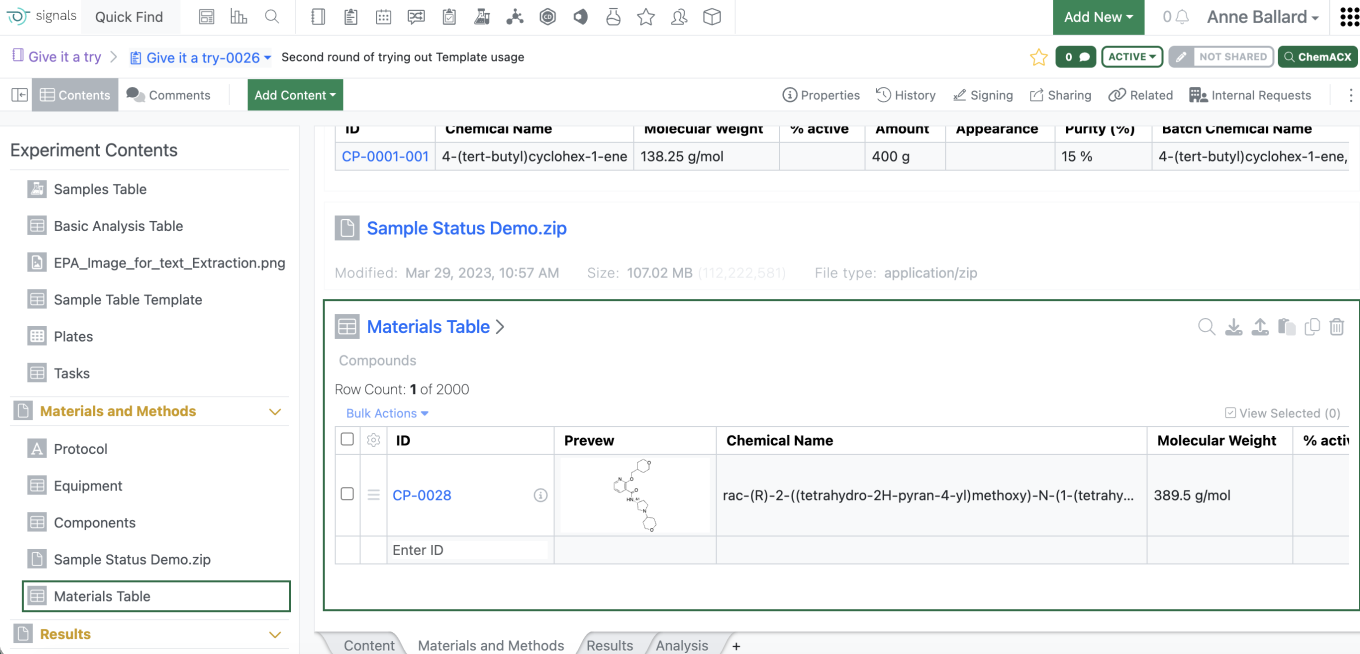
For the end users, if the preview is available, they will be able to see the preview of the structure similar to what can be seen when they mouse of the information icon. If they do not have access to the material, they will not see the structure. Similarly, if a structure is not available, the preview will indicate that the structure isn’t available.
End users can now register sequence-based materials such as DNA, RNA, and Proteins and the pre-configured Plasmids library in a materials table and via bulk import by pasting in a sequence. A raw sequence string, FASTA, GenBank, and GenPept entries can be pasted in when registering materials in a materials table:
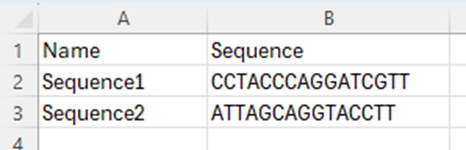

or by including raw sequence strings during a bulk import:
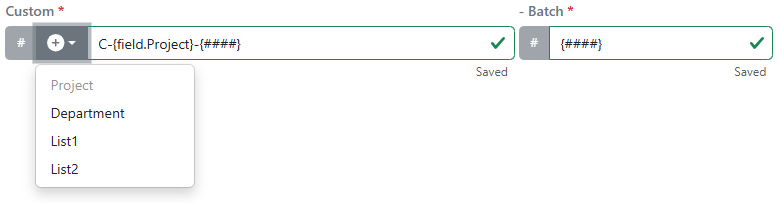
Administrators can now configure asset material IDs to include List and Attribute List field data when registering a material. Up to four configured fields can be included in the asset ID Sequence and will apply to newly registered materials.
Administration

Configuration Transfer for Hierarchical Tables is now available for all systems. The Configuration Report is also now available for all systems.
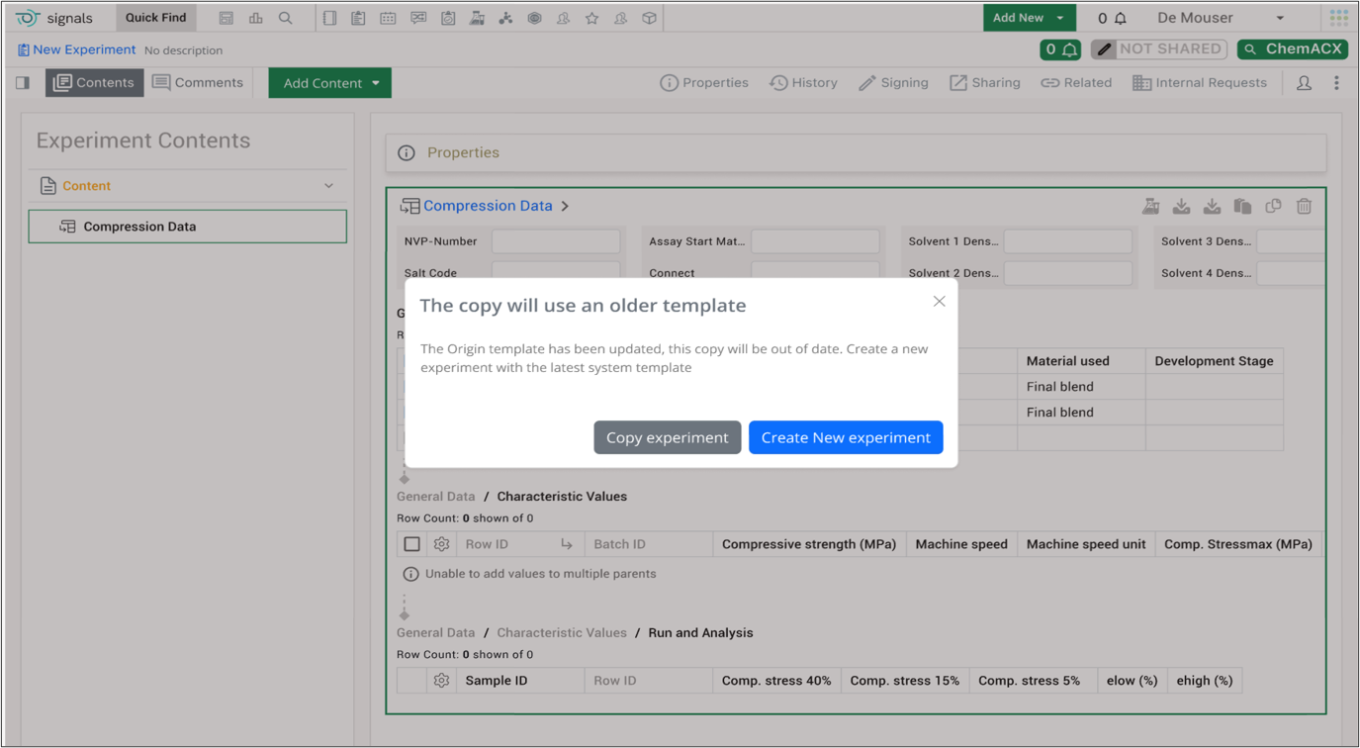
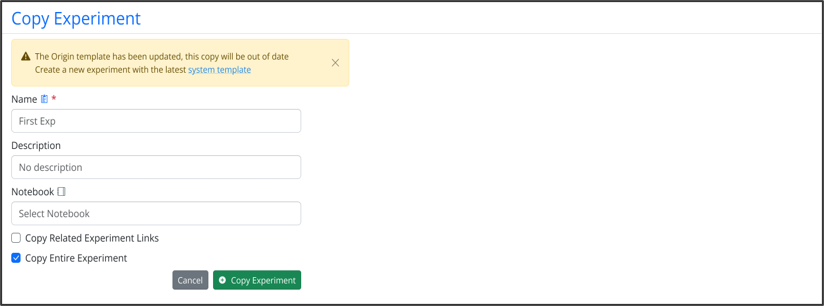
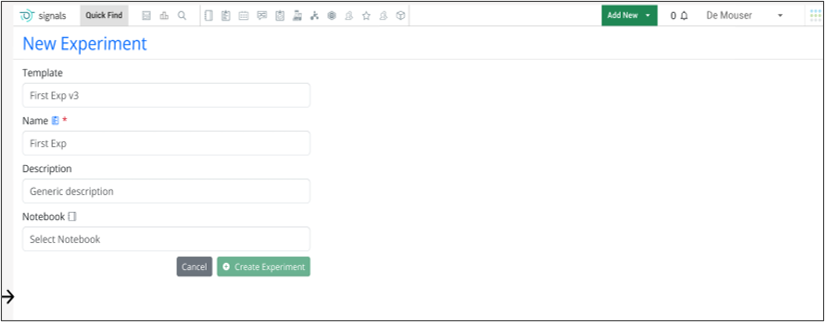
Warning message displays when users clone experiments that are based on outdated system templates. If an admin has updated a template and an end user clones an associated experiment or experiment-like object (i.e. parallel experiment, request, ADO, work order), a warning message explaining the copy is using an older template will trigger. The end user will have the option of proceeding with copying or creating a new experiment based on the updated template.
Integrations & APIs
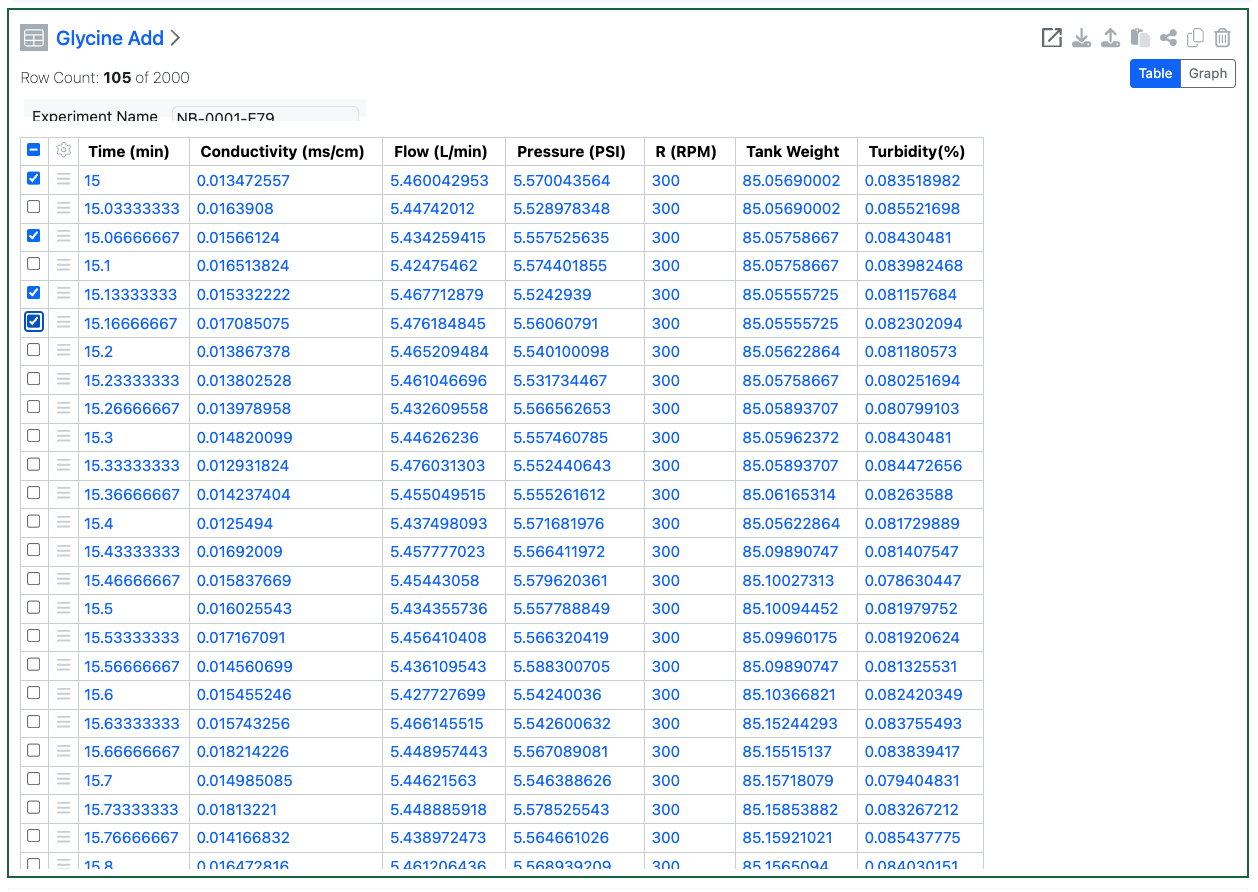
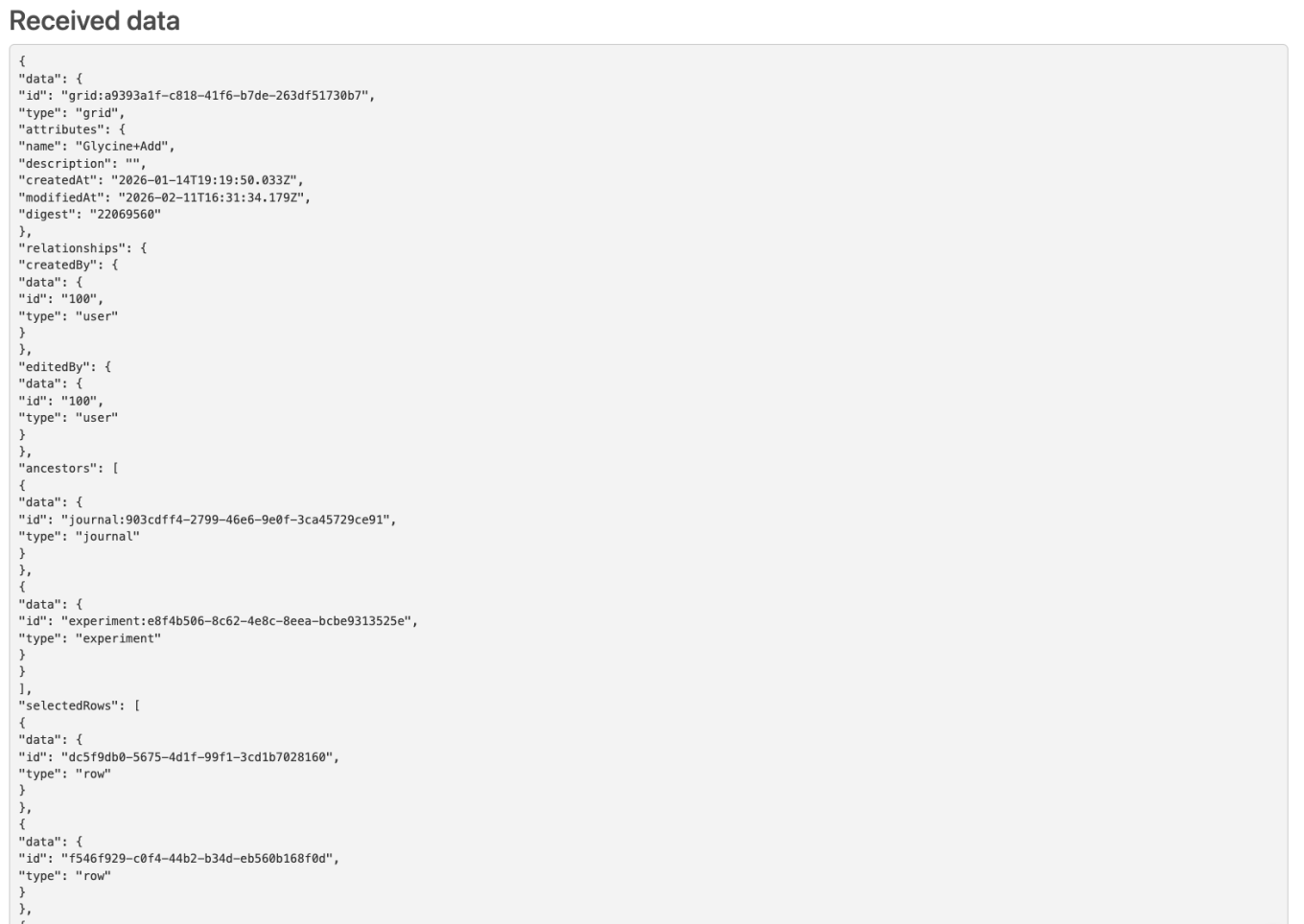
External Actions on Admin Defined Tables and Variation Tables can now be configured to send selected row IDs. If the action is configured with a POST action, the end user will be able to select individual rows in the table. The resulting call will then contain the individual row IDs in the body of the call.
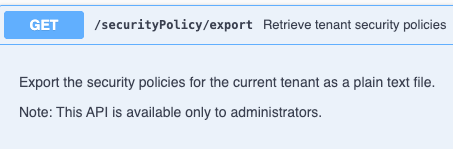
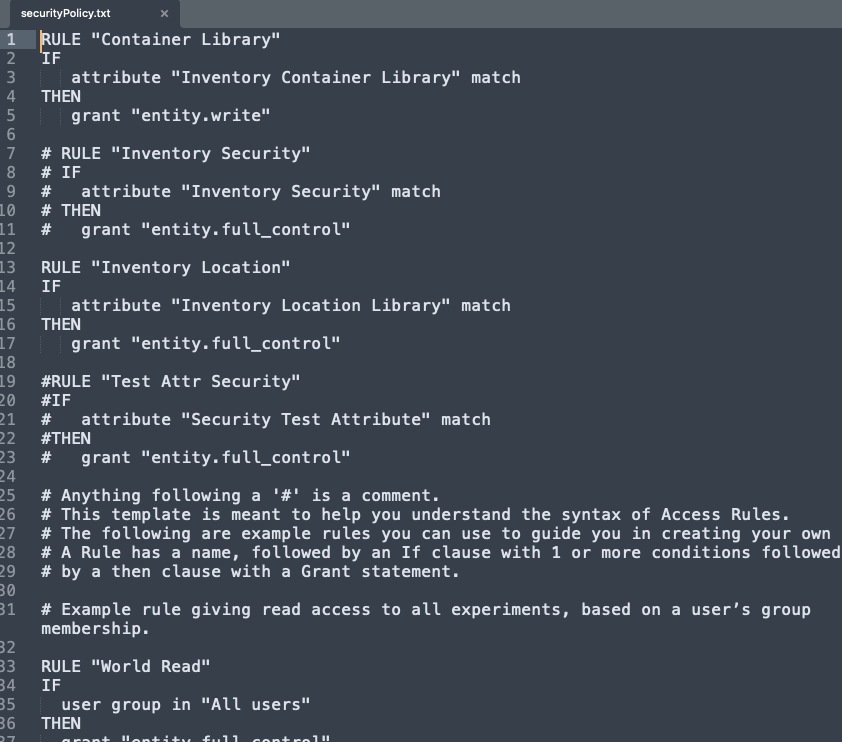
We have added a new API for fetching a text copy of your tenants Security Policy. This API is only available to System Administrators.


We have added enhanced the API for adding Chemical Structures a chemical drawing to support BILN chemical format. The datatype “biln” and the biln string is passed as a in the request body.
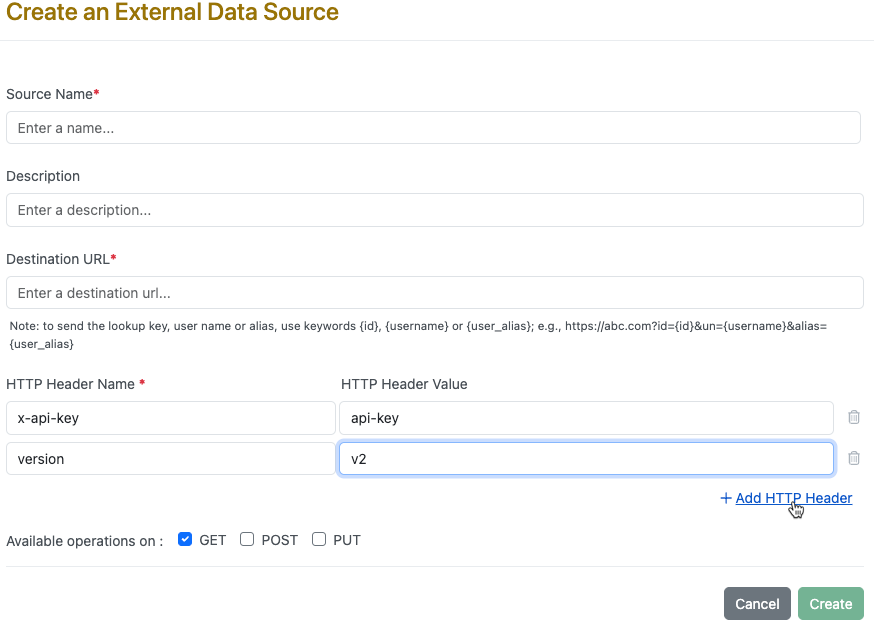
We have enhanced External Notifications, External Data Sources, External Chemical Sources, and External List to allow multiple headers as part of configuration. Existing headers will not be impacted but will allow for additional headers to be added.
Beta Features
The following capabilities are in beta and are available for users, administrators and developers on the Signals platform upon request. Please contact your account representative or our support team if you would like access to the following features.
Fundamentals
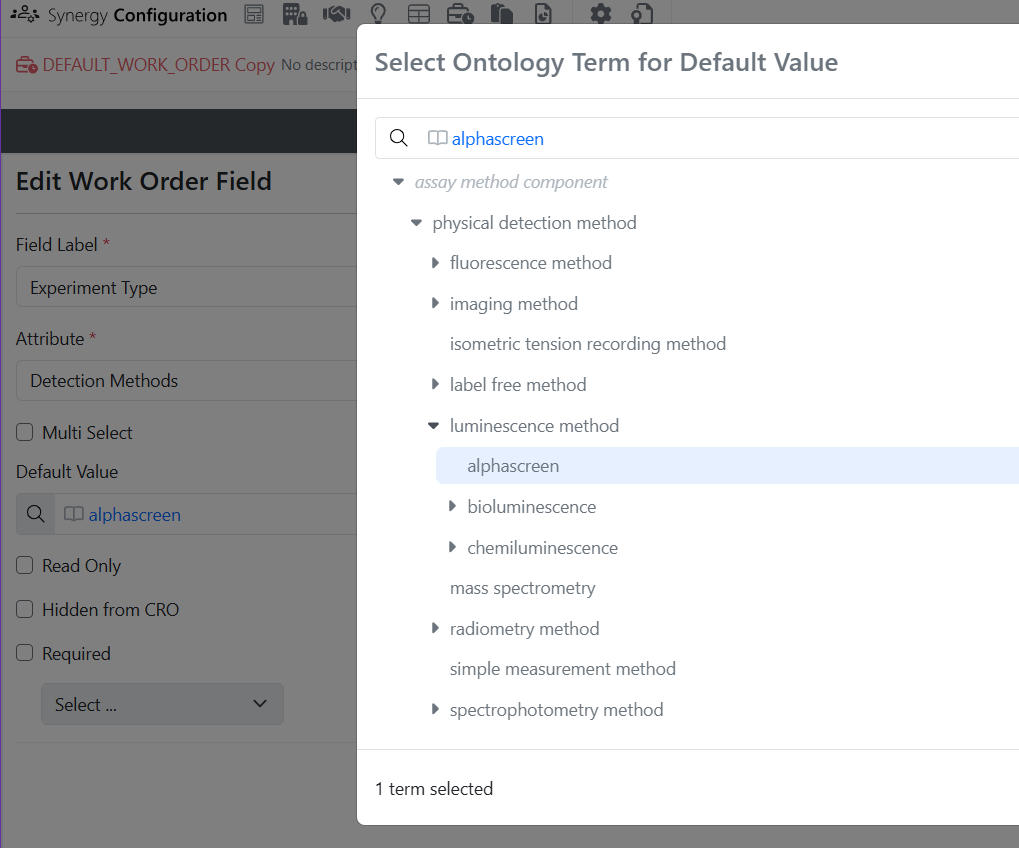
Ontology integration now supported in Synergy, including Collaboration, Idea, Work Order, and Synergy Experiments Templates, as well as Designs and Reference Table fields
Data Workflows & Analytics
Signals Configuration allows access to Analytics definitions.
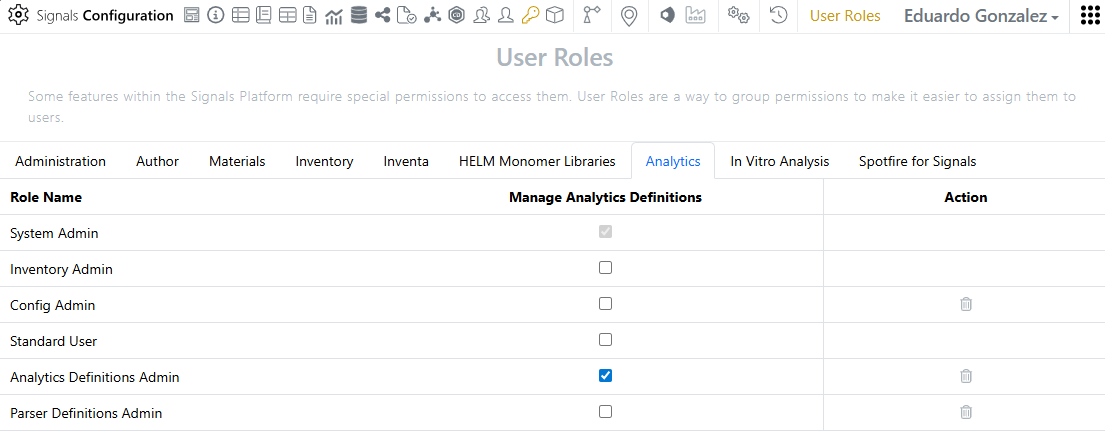
Access to the Analytics definitions is restricted to System administrators and Users with the “Manage Analytics Definitions” privilege.
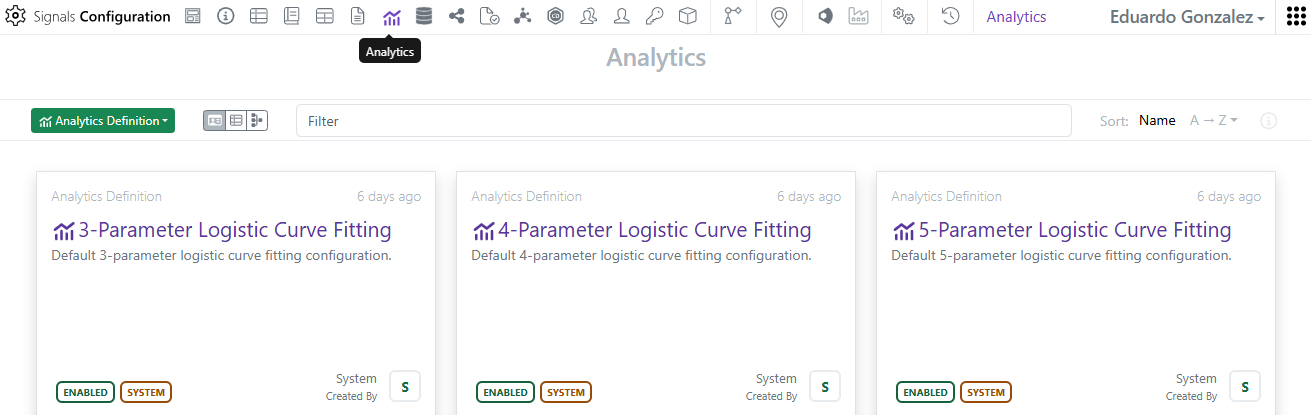
The first set of system Analytics definitions provided out of the box are 3-4-5 Parameter logistic curve fittings.
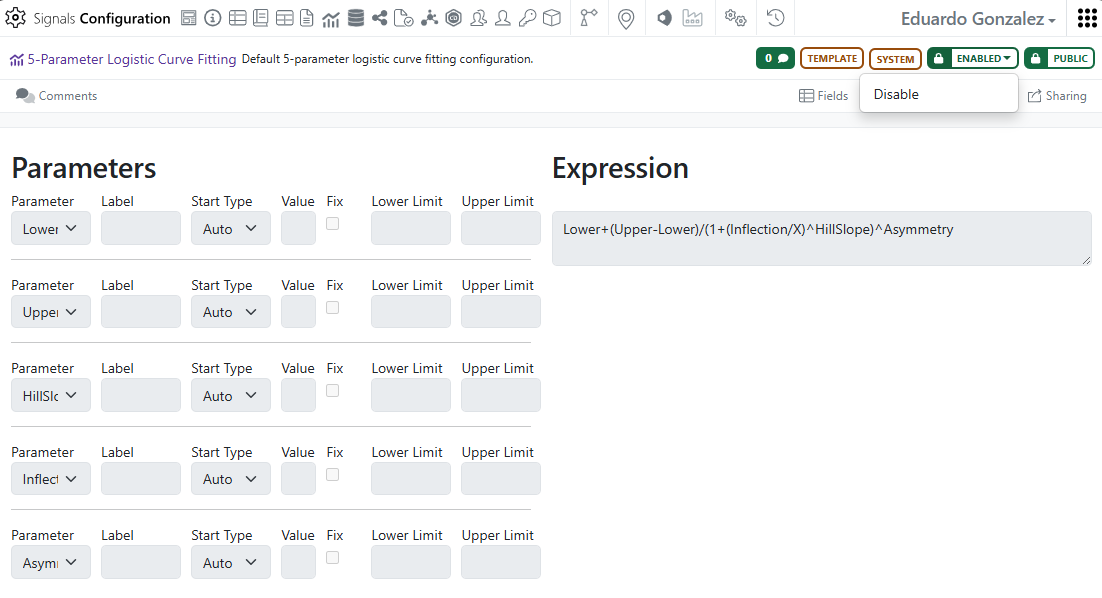
These definitions are currently read-only and can be used in the In Vitro Analysis object, when not disabled.
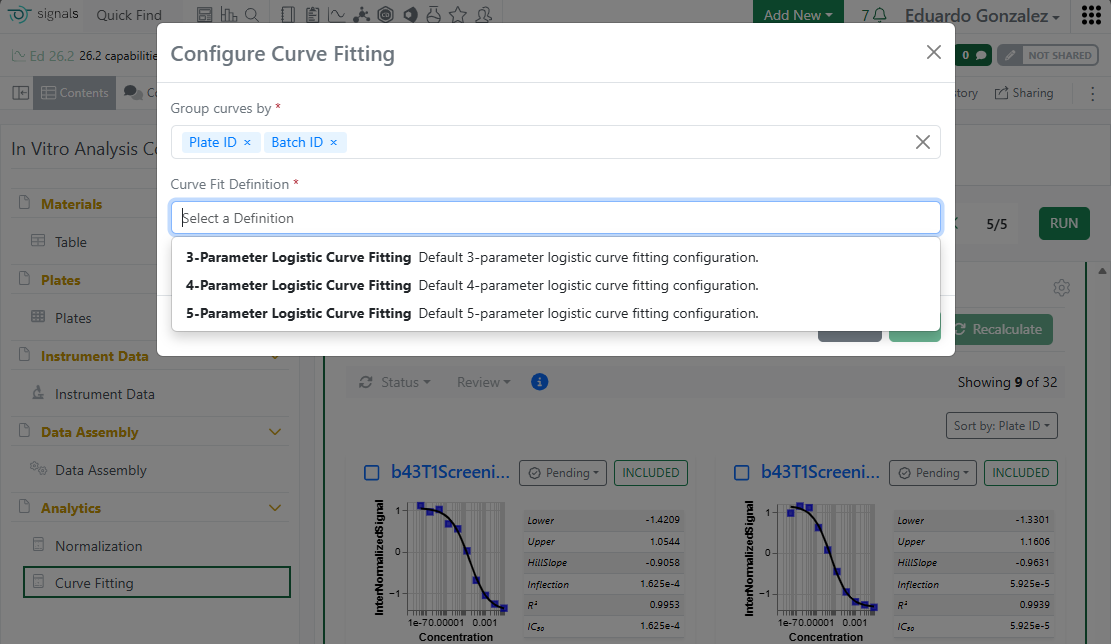
In an In Vitro Analysis, it is now possible to choose any of the 3, 4 or 5 Parameters logistic curve fittings:
Tables
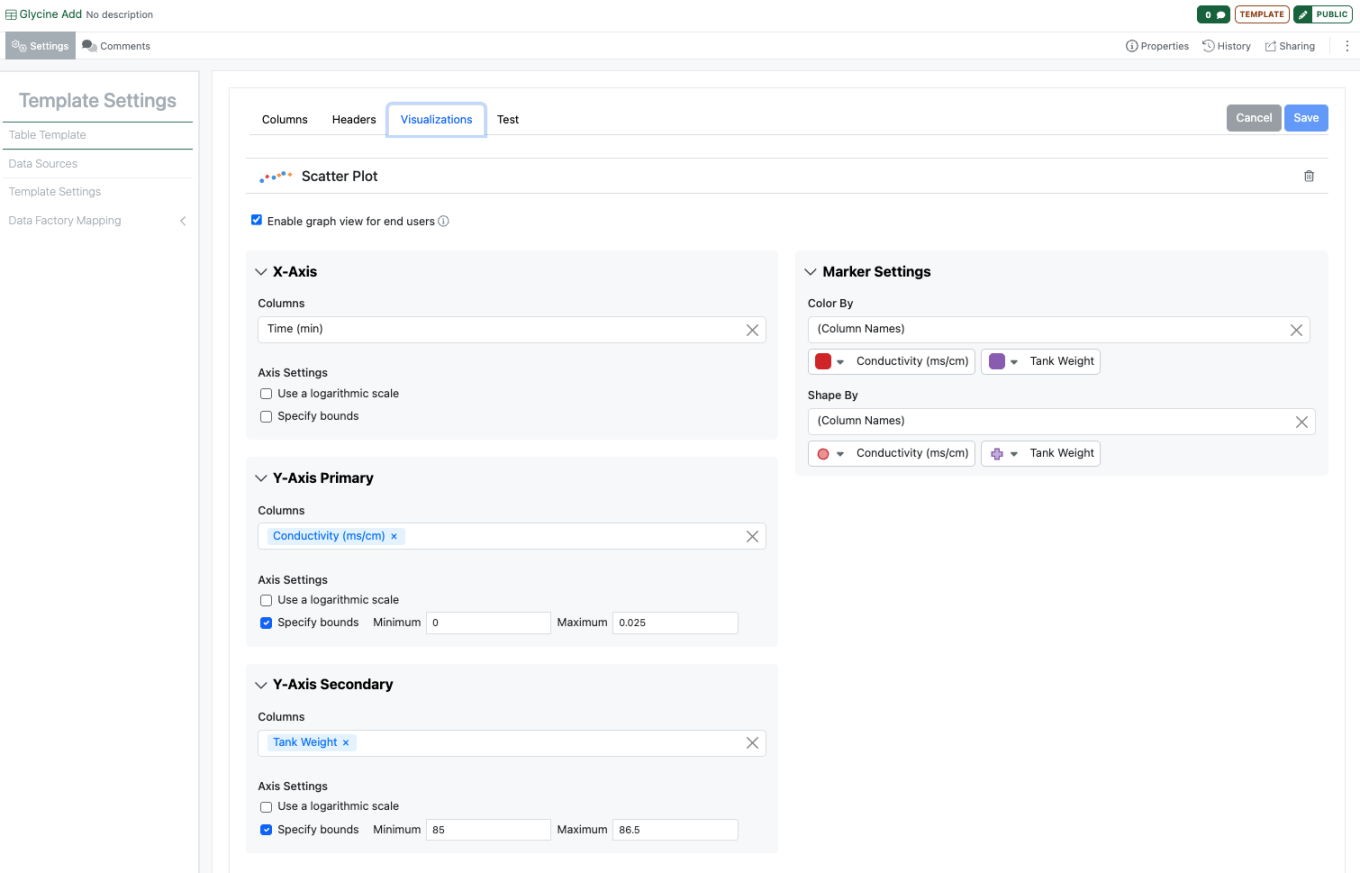
Administrators can now define Scatter Plots that will be displayed in Admin Defined Tables. Administrators can choose if a Scatter Plot is displayed, choose the properties for the axes, the color and shapes and ranges for the axes. If the administrator has representative data they can view a plot via the Try It Out.
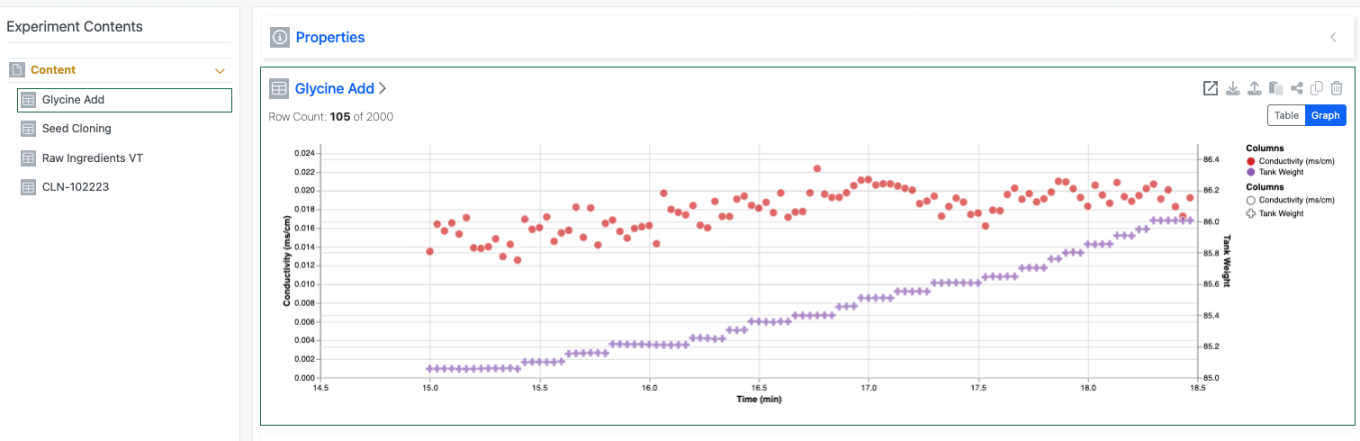
The Scatter plot then appears automatically for the end user once they add data. The end user view is only available to users with a Signals One license.
What's New
We are delighted to release our new version of Signals, 26.1. This release brings enhancements in Tasks, Materials, Data Analytics, Inventory and Tables amongst others. We have new beta capabilities around Ontologies, inherited Material properties and improving the display of tables. The beta capabilities around Contextual Lists are now available to all systems. Finally, we have also fixed a number of small bugs.
The following improvements are available for users, administrators and developers on the Signals platform. Certain features may only be available with appropriate licensing and/or with enablement by an administrator.
- Fundamentals
– Ontologies integration allows terms to be used across experiments, materials, tables, and more (beta) - Data Workflows & Analytics*
– Spotfire-based workflows now integrate the Publish Results to Data Factory app
– Data Assembly download - SAR & Decisioning*
– Bulk Delete published data via the Inventa Dashboard - Chemistry
– HELM editor now supports monomer structure search
– LogP calculation now available in the Analysis panel
– Drag-and-drop of SD/RD files onto the ChemDraw canvas converts to CDXML
– Generate hairpins from oligonucleotides using the auto-pair tool
– Allowing for edits to the Material ID in the Stoich grid
– Updates to Salt Stripping to allow for fractional salts - Tables
– Contextual Lists available to all systems *
– Internal Reference type properties can be used as source for Contextual Lists available to all systems *
– Hidden columns in Tables can be temporarily unhidden
– Calculate Minimum
– Updated admin experience for Admin Defined Tables and Variation Tables (beta)
– Conditional data entry for Number properties in Admin Defined Tables and Variation Tables (beta) - Worksheets
– Contextual List property available to all systems * - Inventory
– Display barcode on Location and Container cards in inventory
– Allow admins to define autofill patterns for grid locations
– Support bulk export of internal requests - Materials
– Type in sequence registration in Materials Library
– Lineage viewer Show/Hide Properties UI update
– Inherited Properties for Materials (beta) - Tasks and Requests
– Updates to the Task Modal to facilitate different modes of task creation - Administration
– Configuration Transfer of Contextual Lists available to all systems
– Configuration Transfer of Contextual Lists in Worksheets available to all systems - Integrations & APIs
– New API to fetch Signals Version
– New APIs to support Hierarchical Tables
– Enhanced API to allow adding Assets to materials table
– New CRO APIs for file attachment and chemical compliance
– Enhanced APIs for bulk updating Containers
* Requires Signals One license
† Requires Signals Synergy license
We also fixed several small bugs in this release. Details of the enhancements are described below.
Administrators should be aware that the Normalized Compound Salt Stripping tool has now been updated to better handle fractional coefficients by altering how multi-fragment salts are interpreted from the Salt List. This change applies to registrations subsequent to this update, existing compounds will remain with the prior understanding until edited.
Data Factory Administrators should note that the syncing material library may now be changed after deleting any Data Factory projects.
Data Factory Administrators should note that legacy Data Factory projects will be fully deprecated in a future release. A straightforward migration path will be provided in an upcoming release to support this transition.
Data Factory administrators should be aware that the behavior of System Generated IDs will be updated in a future release. File-based publications will no longer support System Generated IDs.
Data Factory administrators should also note that the Advanced Security policies in Inventa apply an "AND" logic across all user groups, restricting access if any groups lack permissions. This behavior differs from the standard Signals security model and will be fixed to align with the standard in a future release.
Administrators should be aware that we will be retiring the CRAIS Checker capability from Signals Notebook and Signals One. This capability, which was only available via request, was built to allow integration specifically to the Patcore CRAIS Checker database. Such workflows have been superseded by the Chemical "External Checking" capability which can be used against any chemical compliance database. Any customers still using the CRAIS checker are encouraged to update their workflows to use this capability instead. We encourage any customers who require assistance with this transition to contact our support organization and/or engage with our Professional Services team. The CRAIS Checker capability will be retired by March 31st 2026.
Administrators are recommended to subscribe to the channels within our support news site found at https://support.revvitysignals.com/hc/en-us/categories/360004446171-Support-News which contains more information about releases and other pertinent product information.
This content is anticipated for release on our Production E3 environments, and for Private Cloud customers on our deferred release schedule, in July 2026.
Further Details
The following improvements are available for users, administrators and developers on the Signals platform. Certain features may only be available with appropriate licensing and/or with enablement by an administrator.
Data Workflows & Analytics
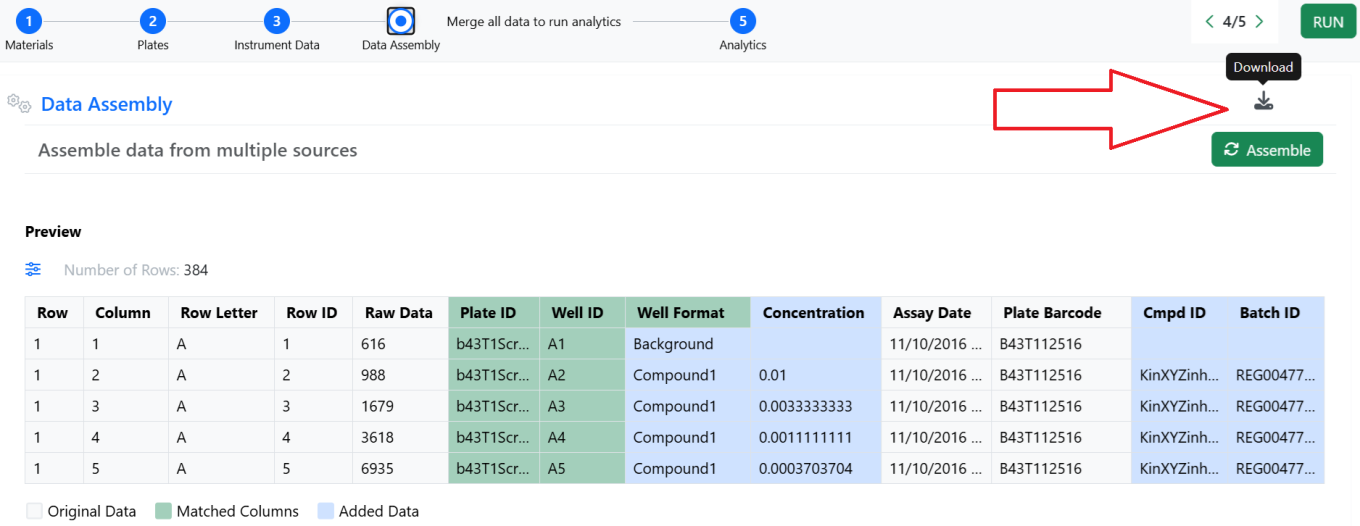
The new app allowing to publish analysis results, associated to mandatory Compound/Batches pairs, to Data Factory 2.0 is now integrated in the out of the box workflows.
The Data Assembly step of the In Vitro Analysis allows assembling the data from the instrument with multiple sources like plate format information and external lookup tables. It is now possible to download, with the click of a button, the full content of the resulting table.
SAR & Decisioning
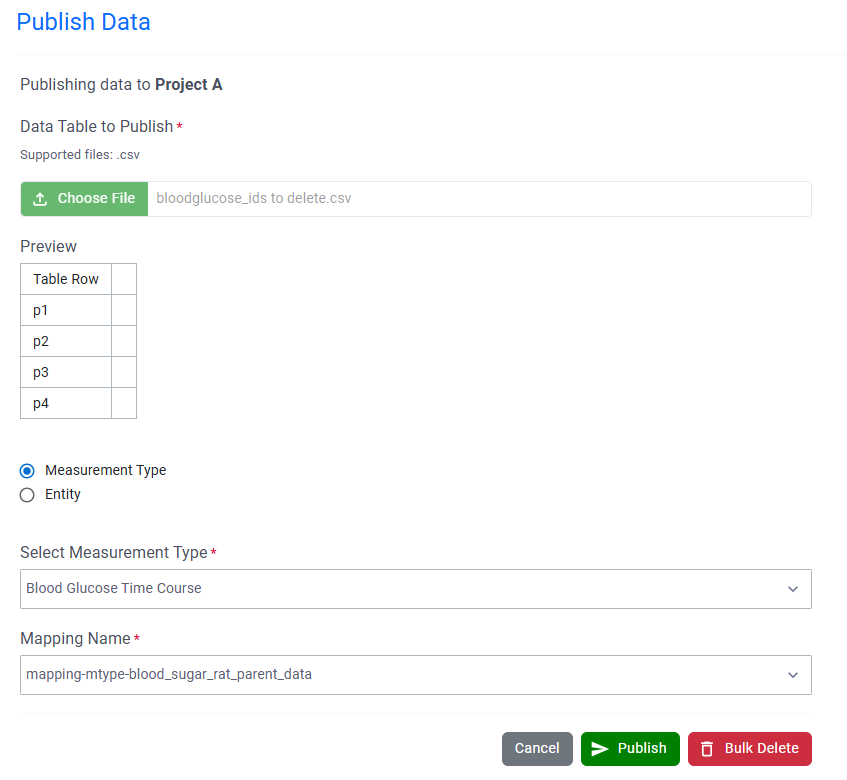
Users can now bulk delete published data via the Inventa Dashboard. After uploading a file that contains a list of Unique Result IDs, users can choose “Bulk Delete” to permanently delete those records from Data Factory. Note: System Generated URIs must be disabled by an Administrator in order to use Bulk Delete.
Chemistry
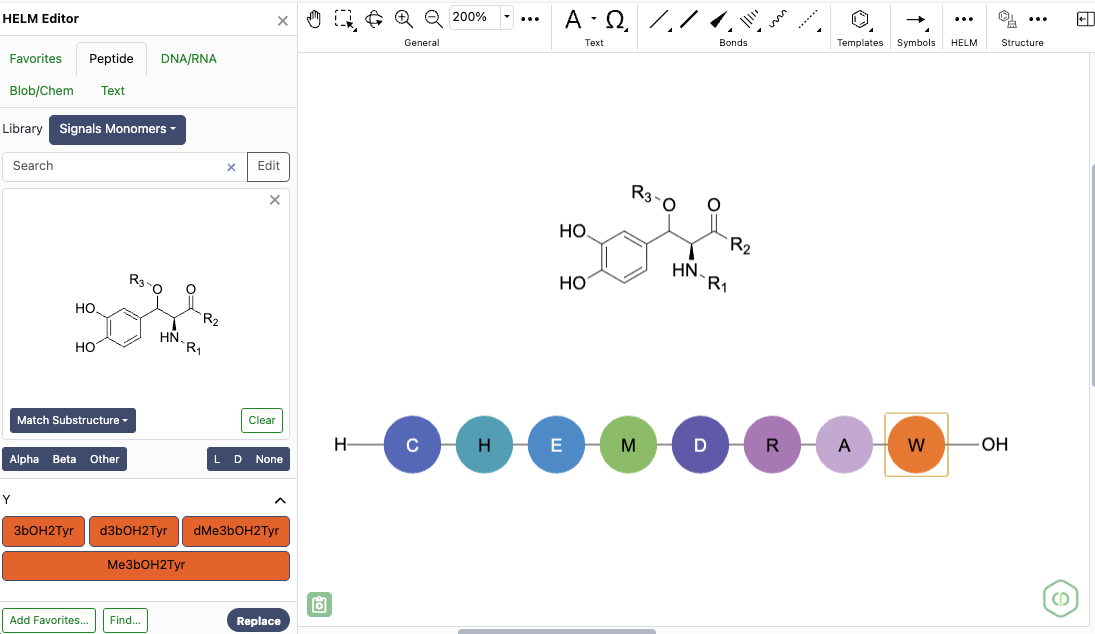
The HELM editor now supports structure-based monomer search alongside text search, using a new 'Structure' button and preview panel in the Peptide, RNA/DNA, and Chem/Blob tabs. Structures can be pasted from common formats (cdxml, cdx, SMILES, molV2000/3000, InChI) and searched as substructure, similar, full, exact, or full including tautomers, with results returned directly in the monomer panel. Structure and text filters can be combined to refine monomer results within the active library.
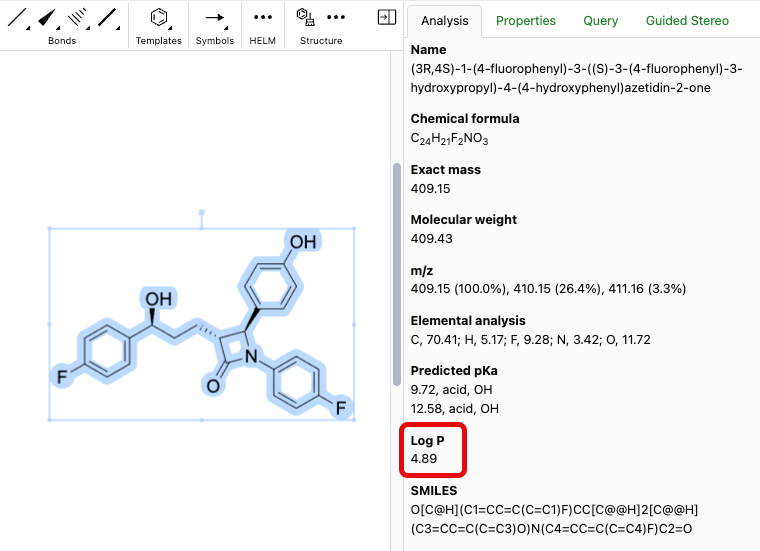
LogP is now available as a calculated property in the analysis panel of the ChemDraw editor. The calculation is based on the current selection and supports structures containing fewer than 1,000 atoms or 150 heavy atoms.
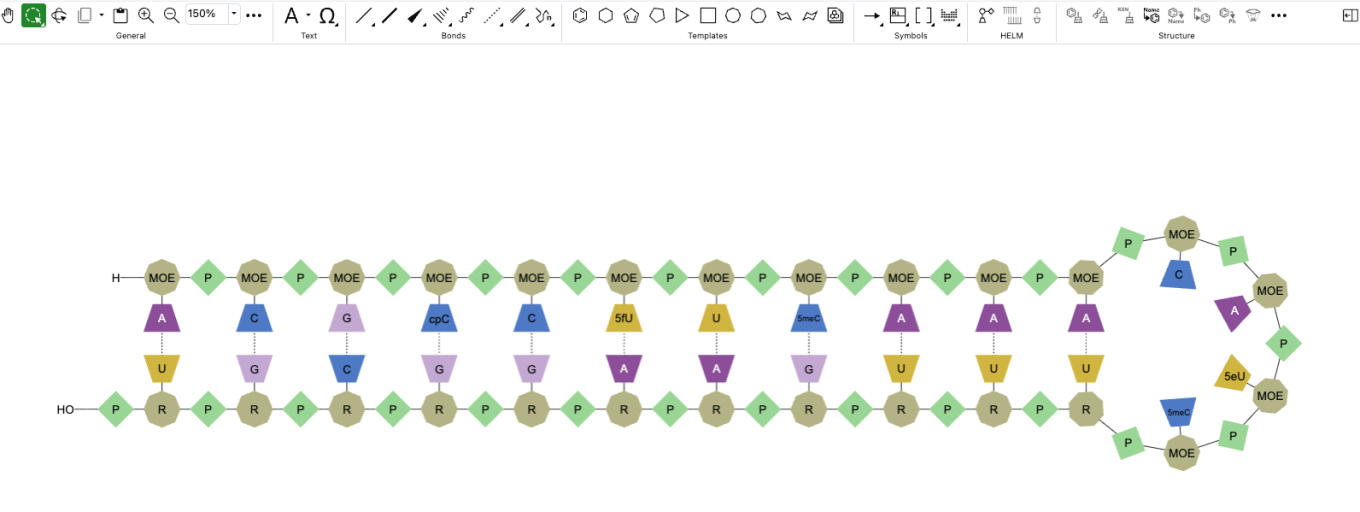
SD and RD files can now be dragged and dropped onto a ChemDraw canvas and converted automatically to cdxml for continued editing.
The ‘auto-pair’ tool in the ChemDraw editor can now generate hairpins from a single oligonucleotide sequence. It identifies the region of maximum complementarity within the strand, places hydrogen bonds between complementary nucleobases, and forms a hairpin loop from the remaining unpaired nucleobases.
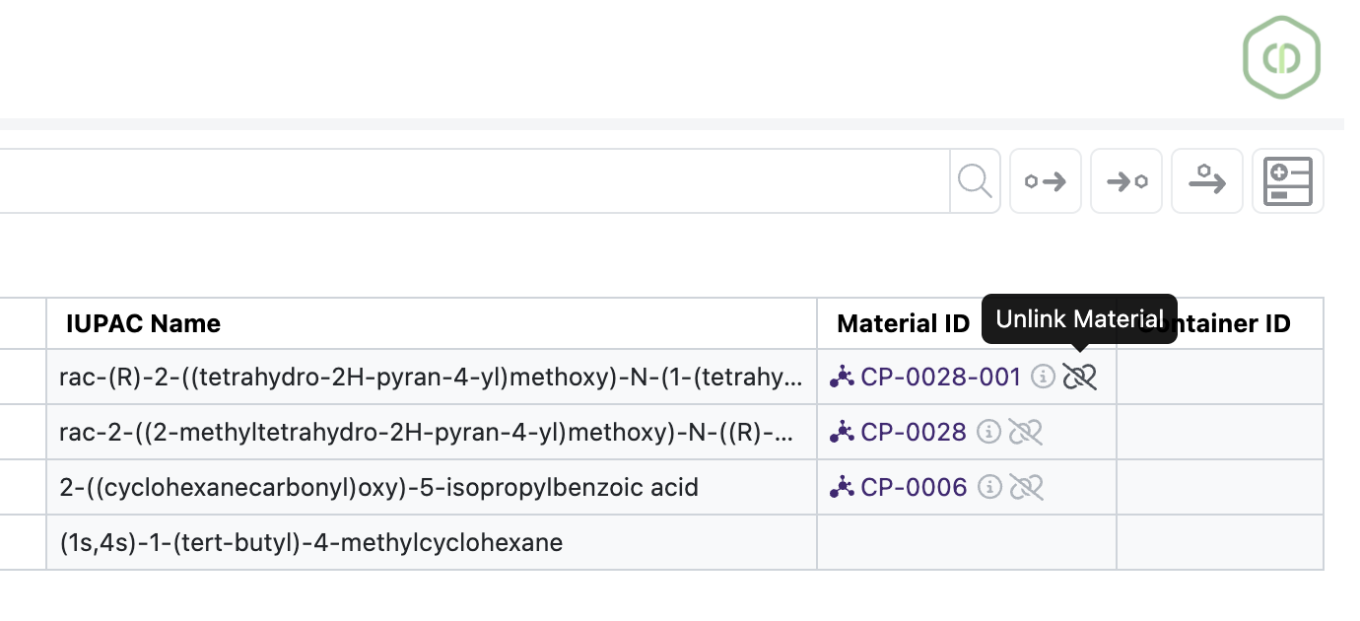
Users will now be able to unlink the Material ID of chemicals added via the Quick Add. Unlinking the material does not erase the chemical from the drawing. It just removes the Material ID which previously could not be removed or altered.
In an update to the behavior of the Salt Stripper for Normalized Compounds, partial salts will now be fully stripped where previously only whole salt coefficients were recognized. This will not alter the batch level chemical name, MF, or mass values as those are calculated from the unaltered batch structure. This will just allow the salt table to more accurately reflect the stripped salts
Tables
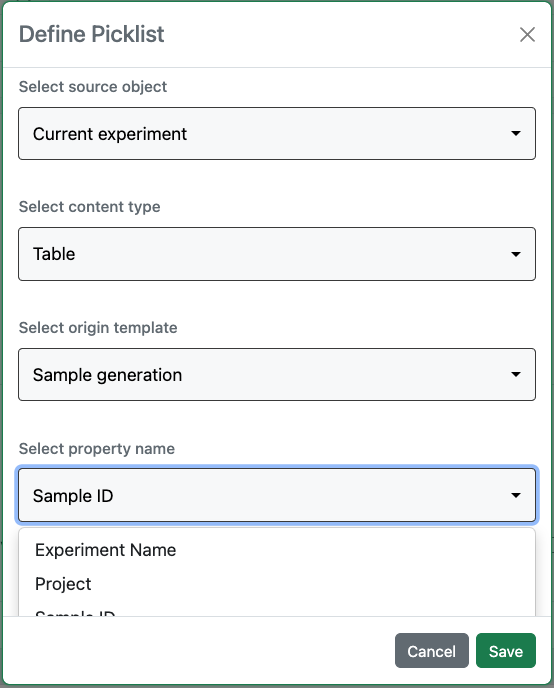
Contextual Lists are now available to systems with a Signals One license.
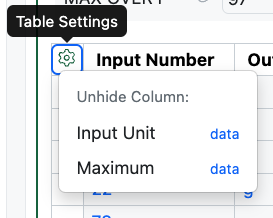
End users can now temporarily display hidden columns in relevant Tables even if they do not have Edit on the content the Table is in. The Cog icon will now be shown and will display in blue if hidden columns contain data. The hidden columns can be exposed from this menu. The display of the columns is transient and the table will be returned to its original state upon a page refresh. This applies to all relevant tables including Admin Defined Tables, Variation Tables, Hierarchical Tables, Design Tables, Sample Tables and Material Tables.
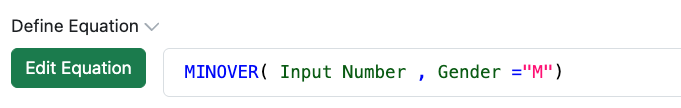
A Configuration Administrator can also define equations to calculate the minimum of a series of data. The functions MIN and MINOVER allow the selection of the lowest number of a set of data. If applied in a Header, the conditional minimum can be calculated using MINOVER or by nesting an IF functions within MIN.
Worksheets
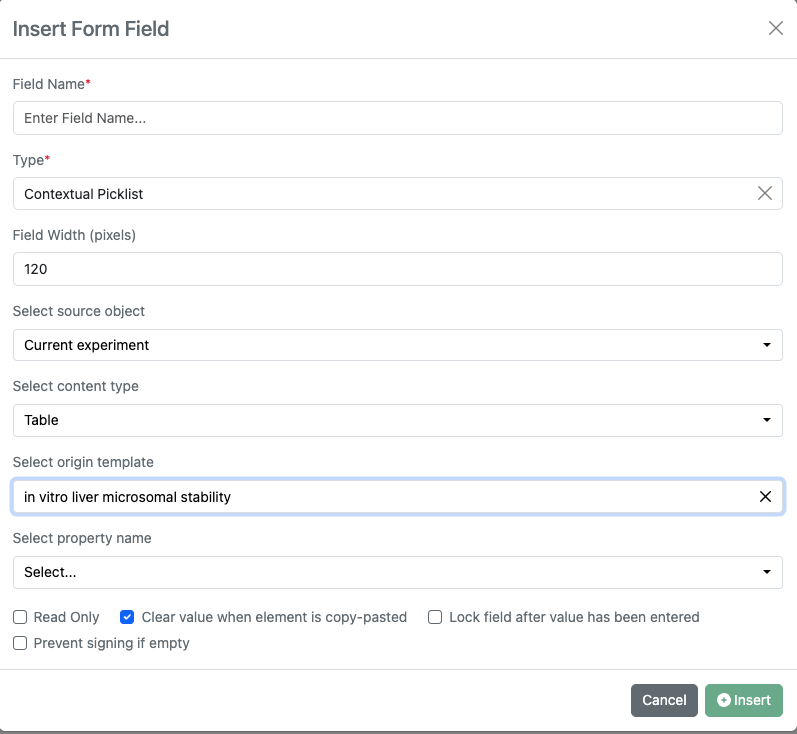
Contextual List properties into Worksheets is now available to all systems with a Signals One license.
Inventory
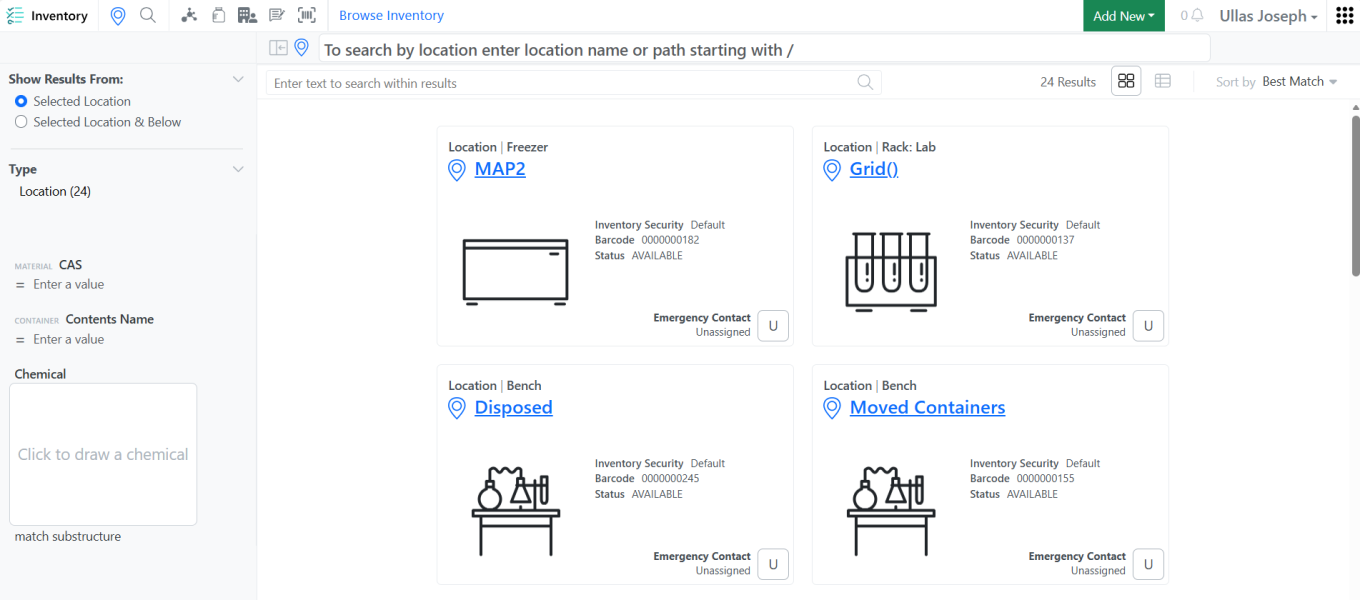
Barcodes are now visible on both the Location and Container Card as well as in the List view within Inventory. The display order of barcodes follows the property sequence defined in Location Type and Container Type settings.
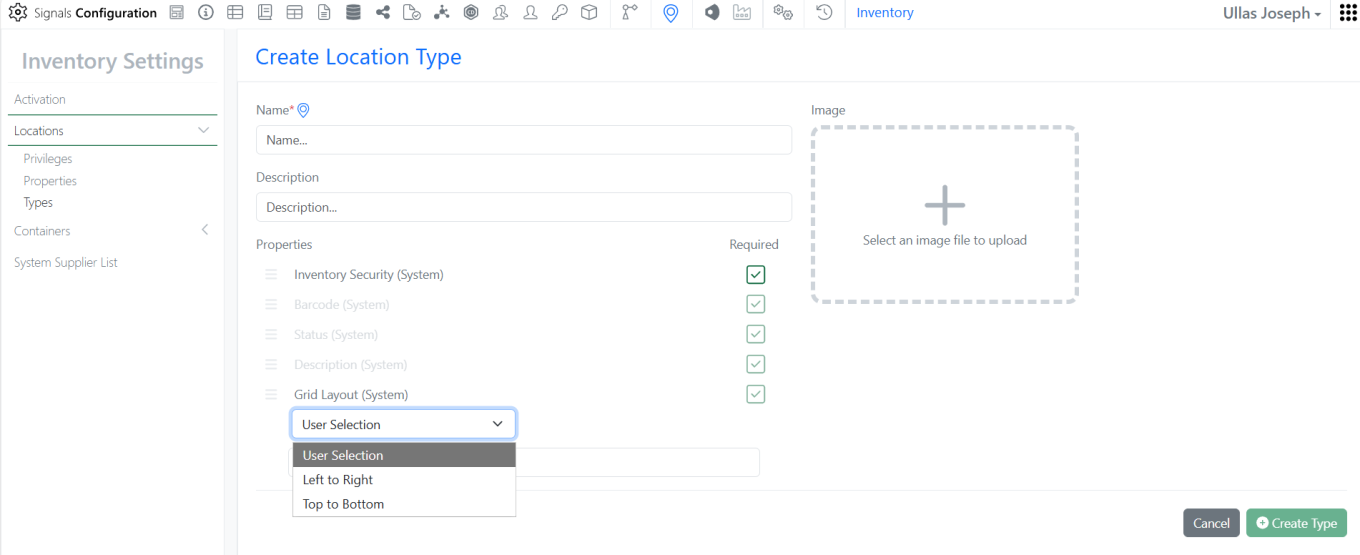
Admin users can now define grid autofill behaviour by setting up pattern mappings for Location Types. Available options include User Selection, Left-to-Right Pattern or Top-to-Bottom Pattern.
This enhancement ensures grid locations are populated according to the pattern selection configured in the Location Type settings.
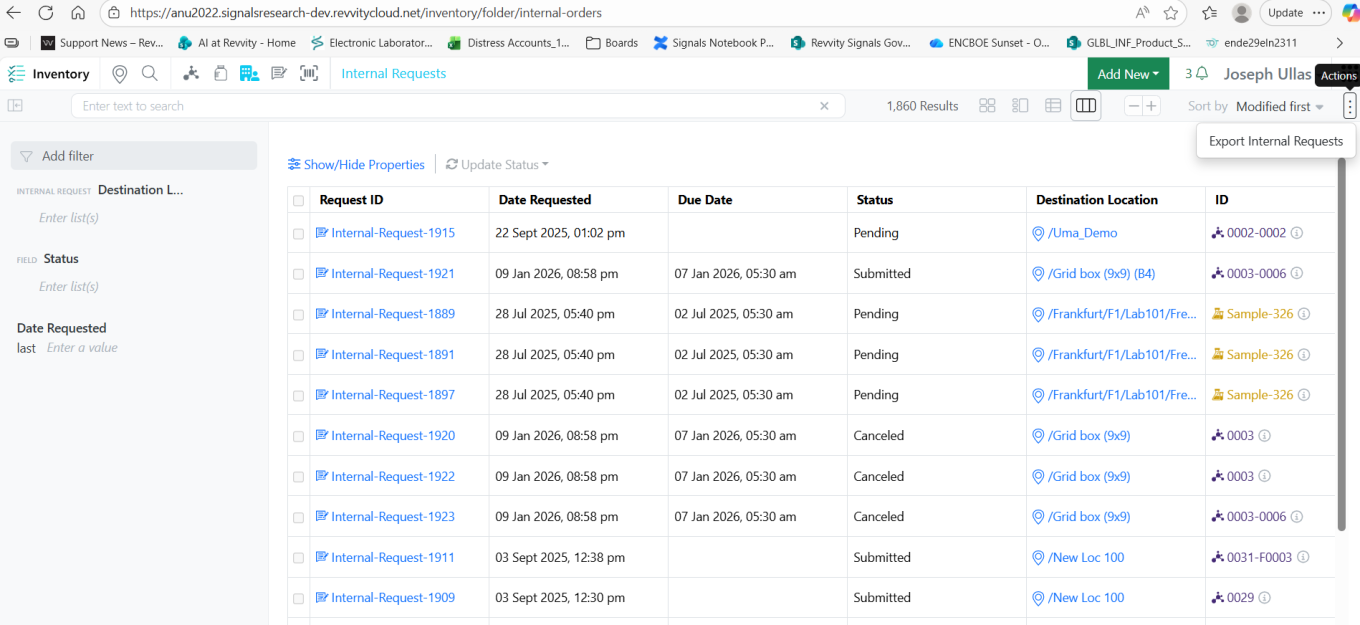
Inventory users with Export Container permissions can now export Internal Requests in two formats, No Structure (CSV) or SDF.
This enhancement allows users to download and share data in the required structure for further processing or integration with other systems.
Materials

End users can now register sequence-based materials such as DNA, RNA, and Proteins and the pre-configured Plasmids library by copying in a sequence from their clipboard. A sequence string, FASTA, GenBank, and GenPept entries can be pasted in when registering materials in the smart folder.
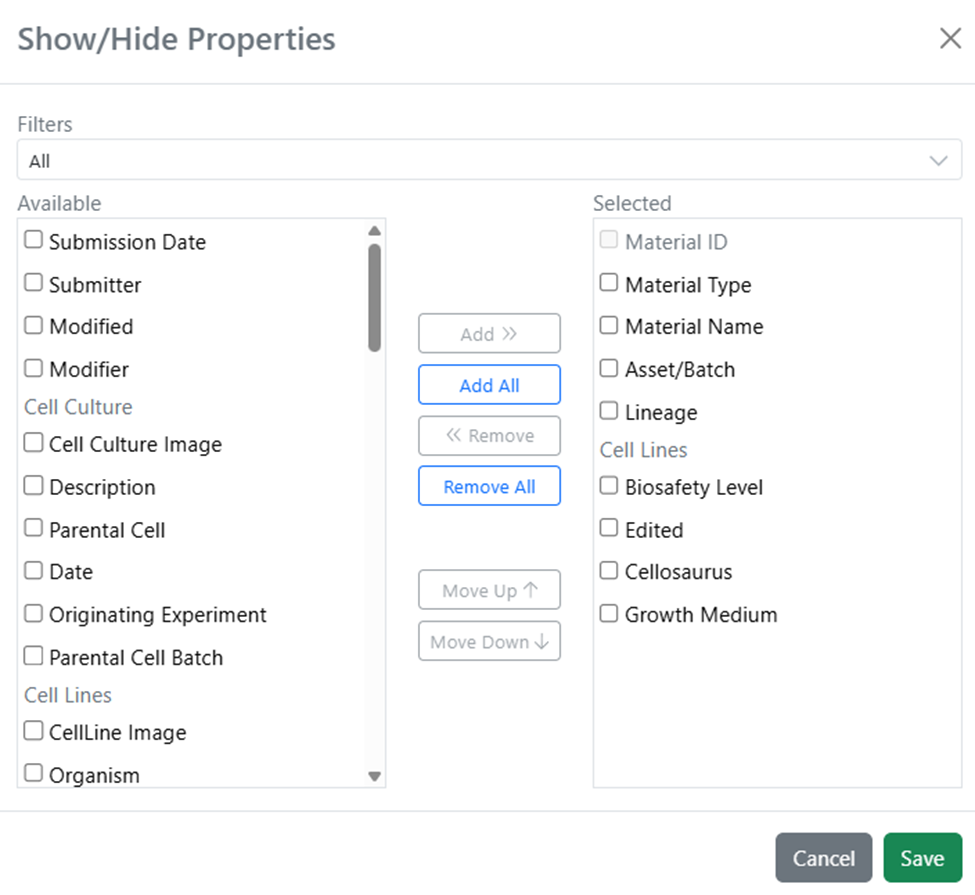
The material lineage table Show/Hide Properties menu has been updated to more concisely display properties from various material libraries present in the viewer. Material library names now act as a header when scrolling through the list of selectable properties.
Tasks and Requests
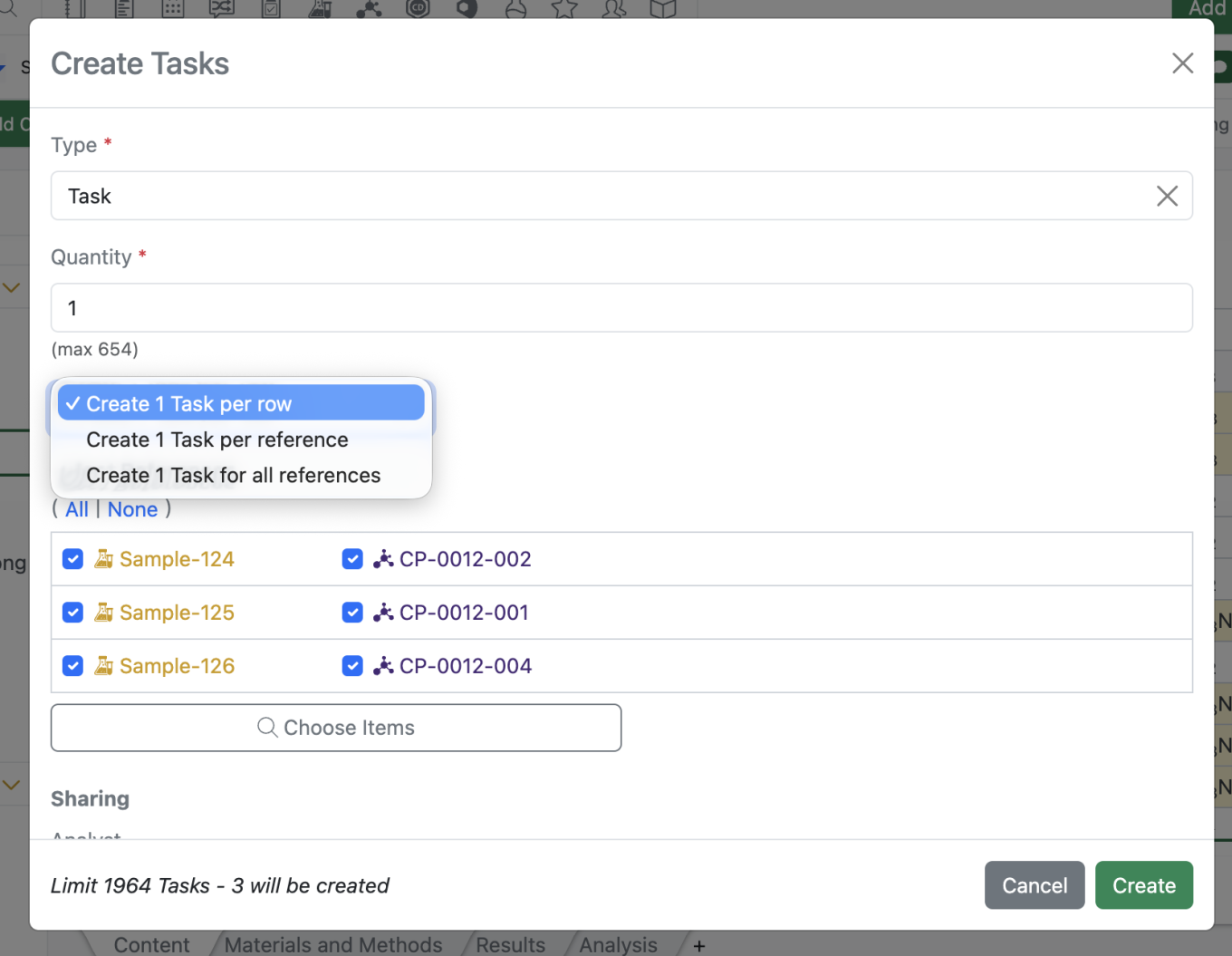
The Task creation modal has been updated to more clearly indicate the number of tasks that will be created and allow for a single task to include all references from the Sample Bulk actions as seen below.
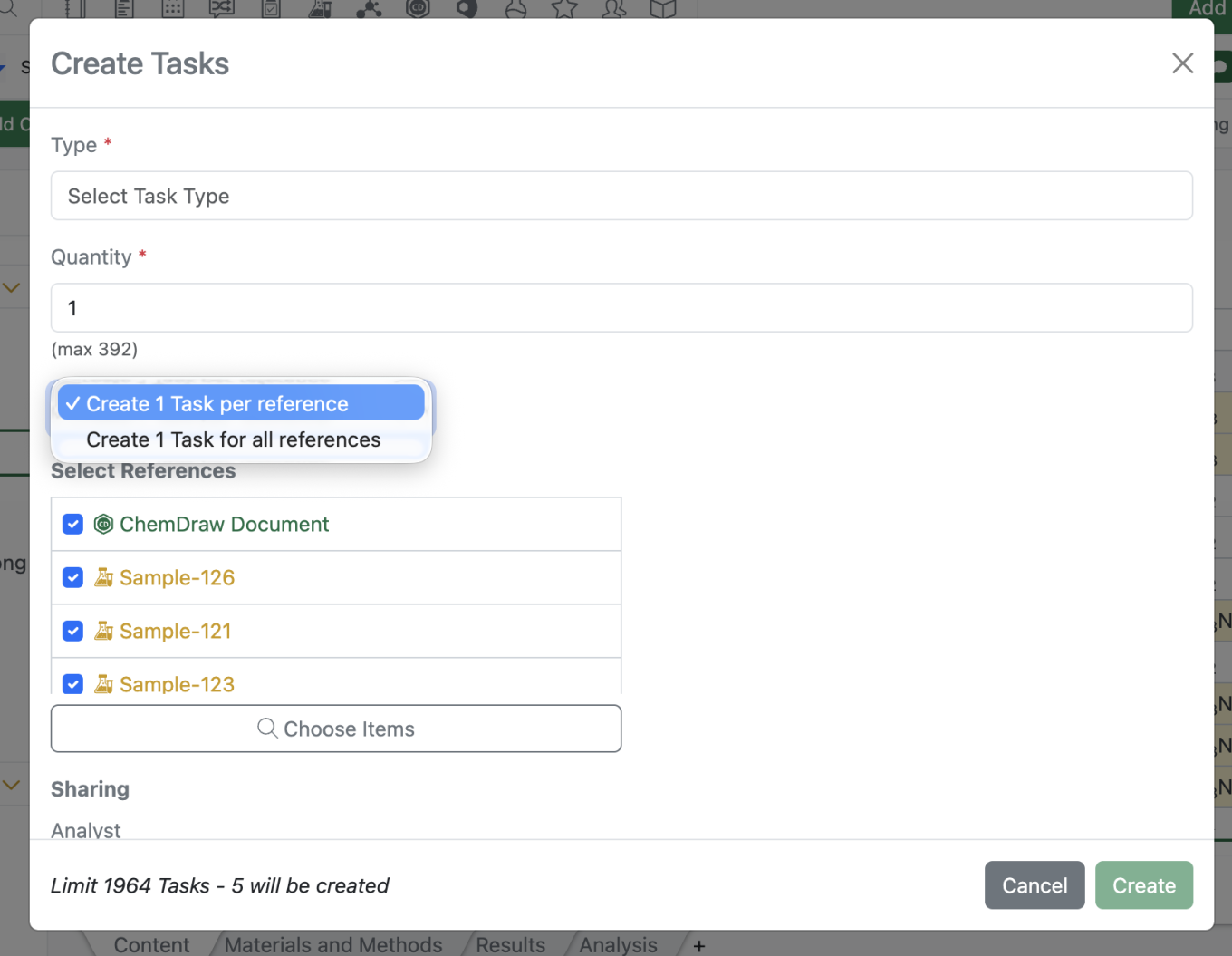
The Task creation modal driven from the “Add Content” menu has also been updated for greater parity to task creation from the Sample Table Bulk Actions.
Updates to the helper text at the modal to indicate the total number of tasks that will be created.
Administration
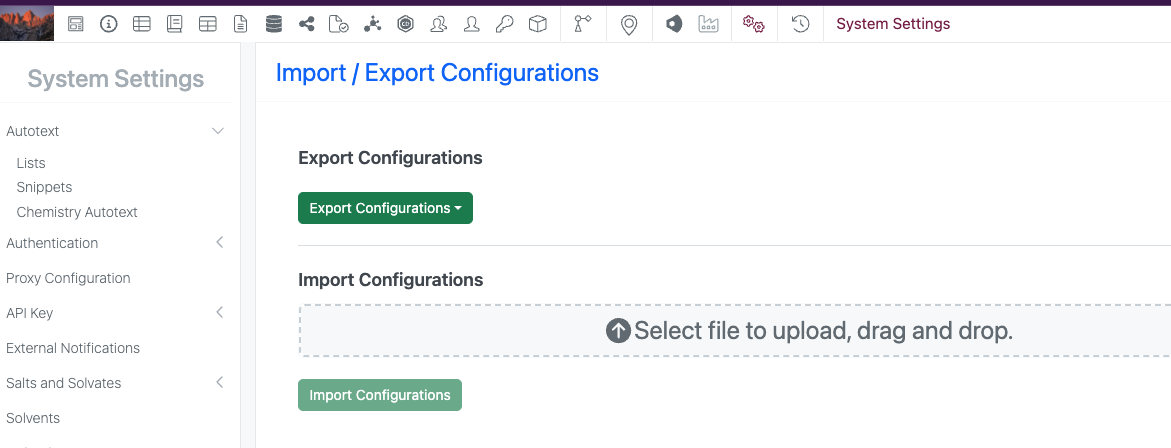
The support for Configuration Transfer of Contextual Lists is now available to all systems with a Signals One license.
Integrations & APIs

We have added a new API for querying your current Signals version.
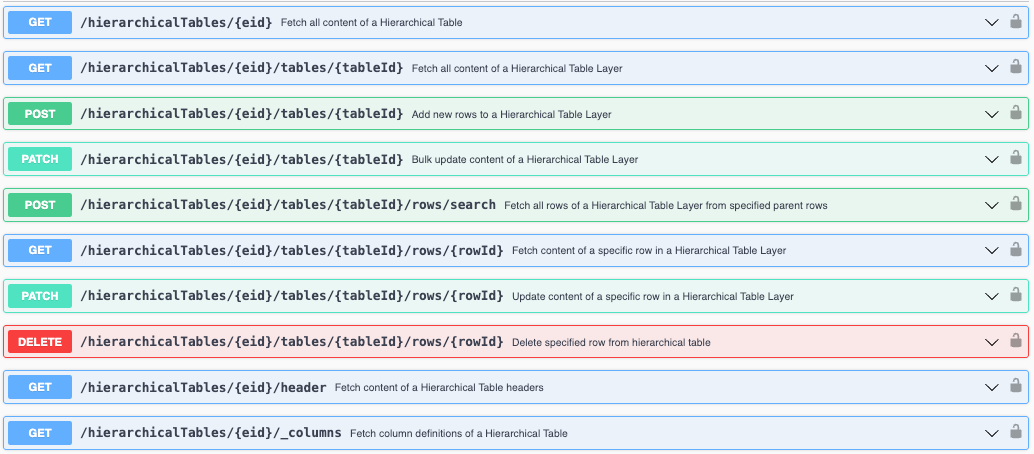

We have added new APIs to support fetching and updating Hierarchical Tables. These APIs allow you to fetch, create, and update content of hierarchical tables.

We have enhanced our API to allow Assets to be added to the Materials Table without requiring a Batch/Lot. The request body will accept the Asset ID where previously it required a Batch ID.
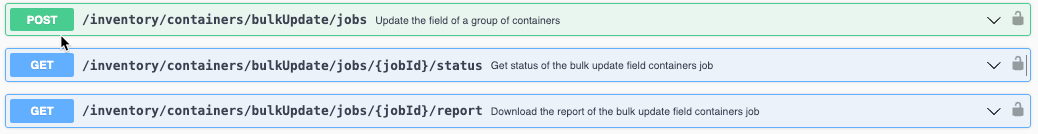
We have added new APIs to bulk update Responsible Person, Amount, and Expiration Date for Containers. This will update the selected field for all Containers included in the request body. Only one of these fields can be bulk updated at a time for all Containers.
The API is asynchronous and kicked off in three steps: Create the Bulk Update Field Job -> Check the Status of the Job -> Get a report of a completed Job as a CSV.
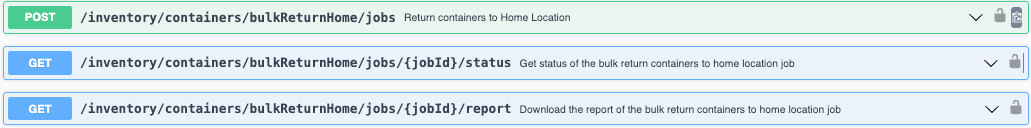
We have added new APIs to bulk return Containers to their Home Locations. This will return all the containers to their Home locations. Only containers with a Home location set will be returned.
The API is asynchronous and kicked off in three steps: Create the Return Home Job -> Check the Status of the Job -> Get a report of a completed Job as a CSV.
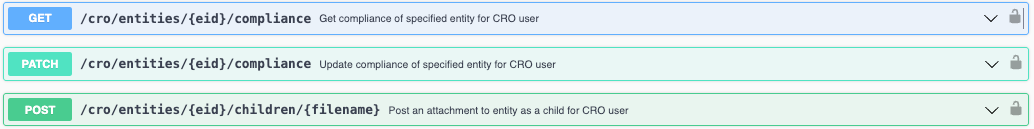
We have added new APIs specific to our Synergy Product CROs for attaching files and fetching/updating chemical compliance. These allow for specific APIs to be utilized by CRO users. This enables customers External Actions to read and write for CRO users without exposing details that CROs are restricted from seeing or interacting with.
Beta Features
The following capabilities are in beta and are available for users, administrators and developers on the Signals platform upon request. Please contact your account representative or our support team if you would like access to the following features.
Fundamentals
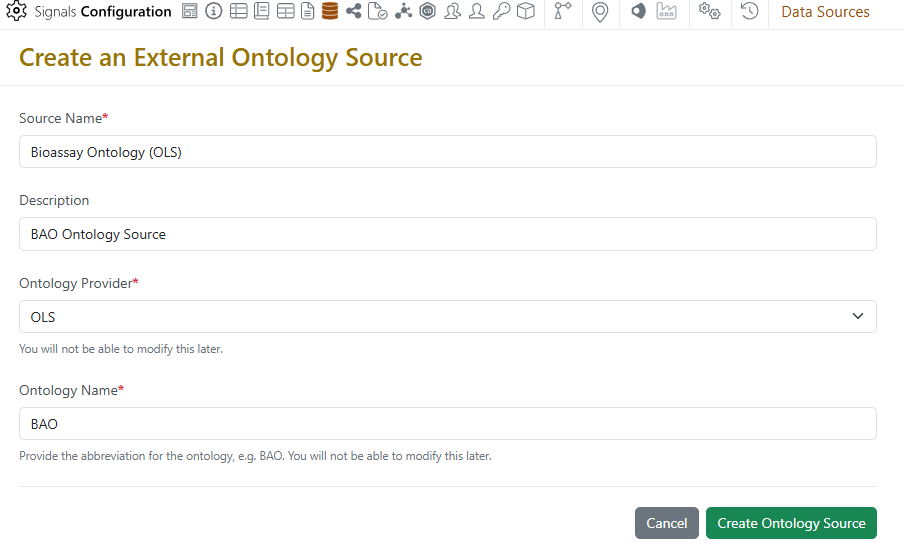
Ontologies are now supported in Signals. Administrators may connect to most public ontologies available on either the BioPortal or Ontology Lookup Service (OLS) providers.
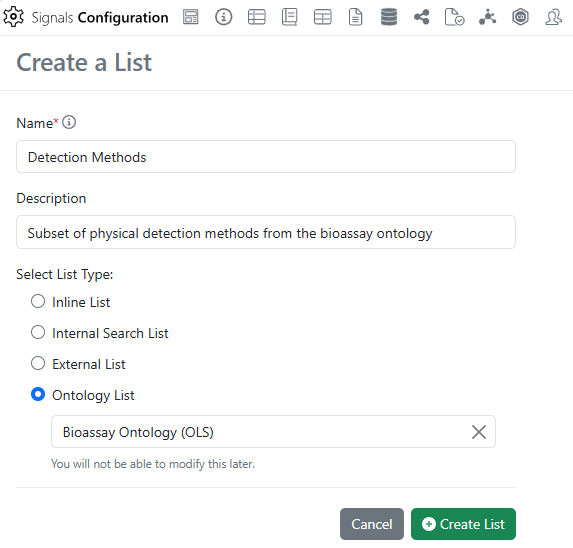
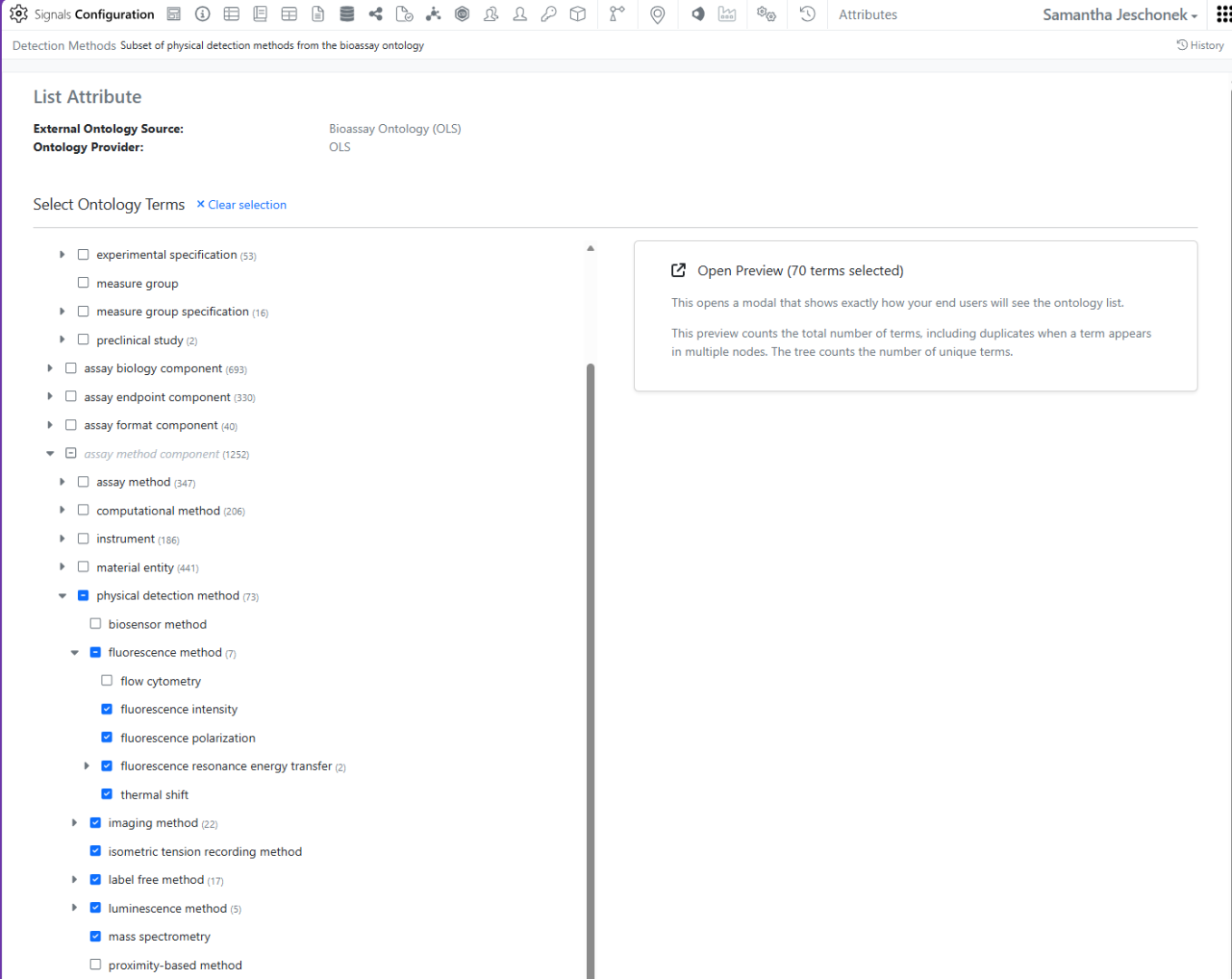
After connecting to an External Ontology Source, Administrators can subset a specific Ontology into a new type of Attribute List called an Ontology List.
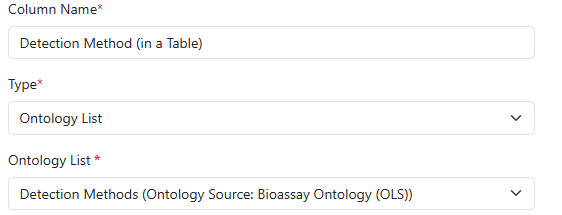
Admins can incorporate Ontology Lists into most Templates within Signals where Attribute Lists are already supported, including Material Definitions, Experiment Templates, Table Templates, Worksheet Templates, Sample Properties, and more
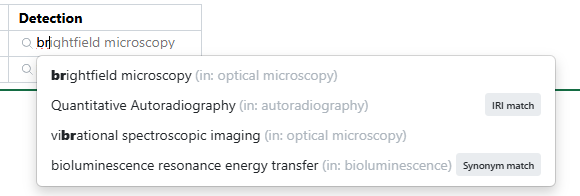
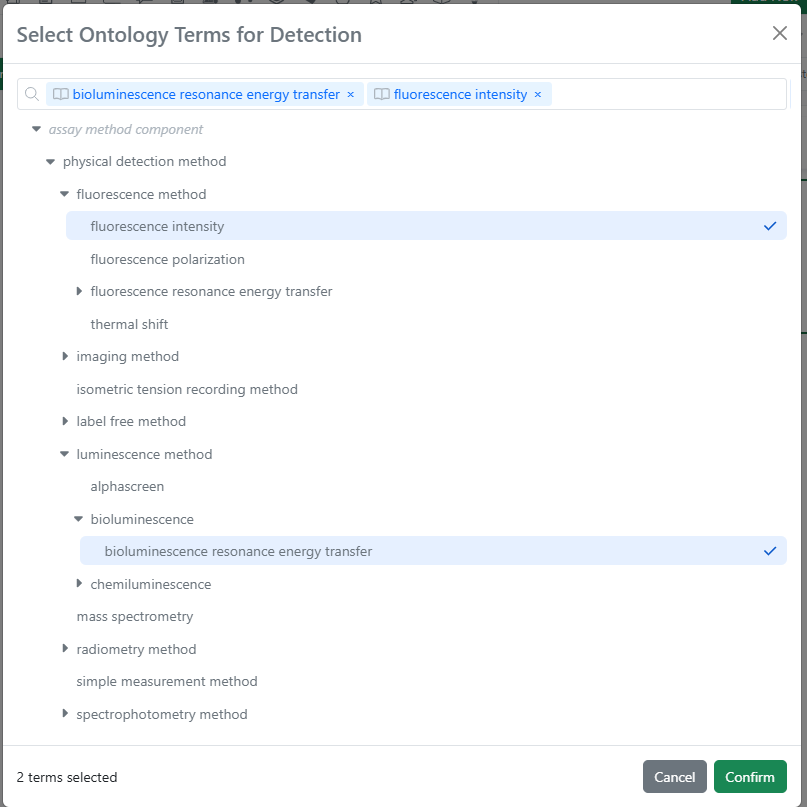
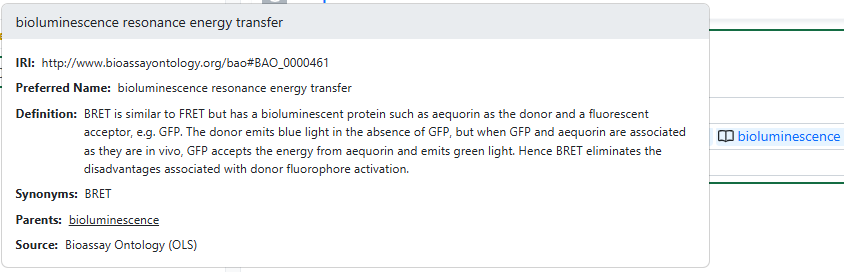
When interacting with a field associated with an Ontology List, users can type-ahead to find a term directly (including matching based on synonym). Alternatively, users can select the magnifying glass icon to browse the hierarchical ontology tree.
You can now split a container into multiple sub-containers with an option to make all aliquot amounts identical. This helps save time and reduce manual entry when preparing multiple identical vials.
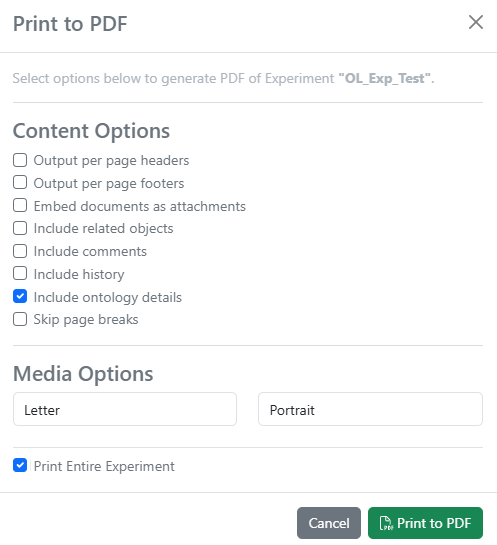
When printing Experiments or other objects to PDF, ontology details can be optionally included, which will embed a CSV with ontology metadata, like Preferred Name and IRI.
Tables
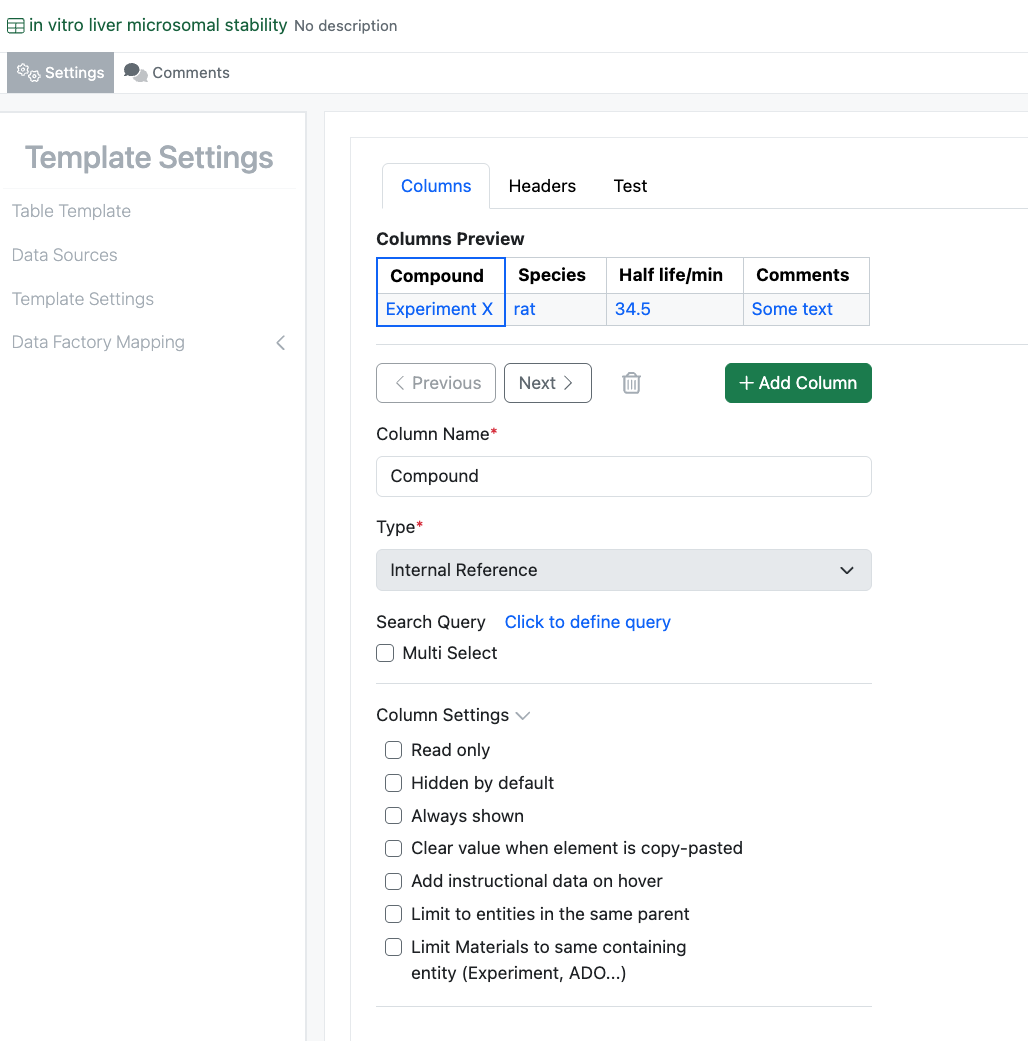
The administrative experience for Admin Defined Tables and Variation Tables has been updated to match Hierarchical Tables. This updated design matches that previously released for Hierarchical tables and provides a smoother experience for defining and reordering properties within the table.
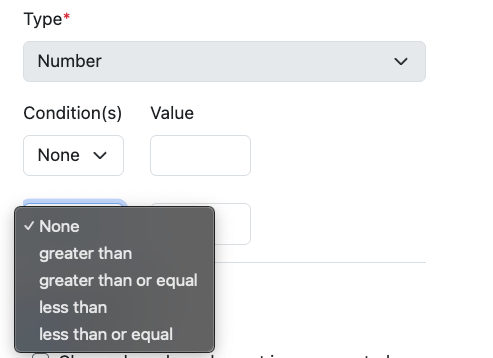
Included within the administrative update described above, we have also extended the configuration of Number properties in Admin Defined Tables and Variation Tables. Number properties can be defined with conditions for data entry, ranges can be set and the property set to either not allow an out of range value, or to provide a visual indication when a value does not match.
Materials
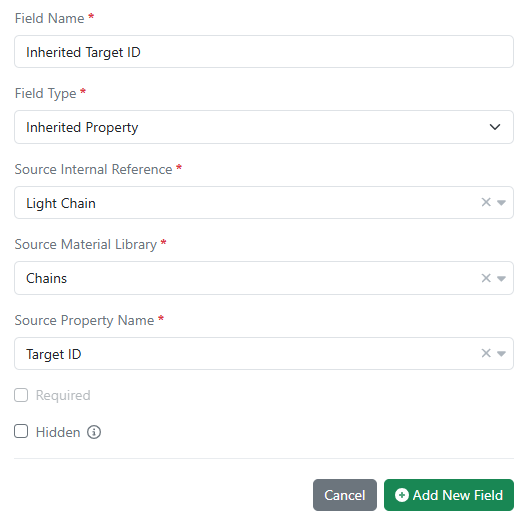
Administrators can now configure inherited properties in all material libraries. Data field values can now be passed down (i.e. inherited) from one material to another using internal reference links. Updates to the source data are automatically propagated to any inheriting entity and are kept in sync.